+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9912 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of alpha2BAR-Gi1 complex | |||||||||
 Map data Map data | alpha2BAR-Gi1 complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / Complex / cryo-EM / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationActivation of the phototransduction cascade / Olfactory Signaling Pathway / negative regulation of uterine smooth muscle contraction / alpha2-adrenergic receptor activity / Adrenaline signalling through Alpha-2 adrenergic receptor / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Prostacyclin signalling through prostacyclin receptor / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / G alpha (z) signalling events ...Activation of the phototransduction cascade / Olfactory Signaling Pathway / negative regulation of uterine smooth muscle contraction / alpha2-adrenergic receptor activity / Adrenaline signalling through Alpha-2 adrenergic receptor / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Prostacyclin signalling through prostacyclin receptor / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / G alpha (z) signalling events / Glucagon-type ligand receptors / G beta:gamma signalling through PI3Kgamma / G beta:gamma signalling through CDC42 / Sensory perception of sweet, bitter, and umami (glutamate) taste / alpha-2C adrenergic receptor binding / epinephrine binding / phospholipase C-activating adrenergic receptor signaling pathway / Adrenaline,noradrenaline inhibits insulin secretion / ADP signalling through P2Y purinoceptor 12 / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / Thromboxane signalling through TP receptor / G beta:gamma signalling through BTK / positive regulation of uterine smooth muscle contraction / Adenylate cyclase inhibitory pathway / Thrombin signalling through proteinase activated receptors (PARs) / alpha-1B adrenergic receptor binding / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / negative regulation of norepinephrine secretion / G alpha (s) signalling events / negative regulation of epinephrine secretion / Ca2+ pathway / G-protein activation / Extra-nuclear estrogen signaling / G alpha (12/13) signalling events / G alpha (q) signalling events / Vasopressin regulates renal water homeostasis via Aquaporins / GPER1 signaling / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / regulation of vascular associated smooth muscle contraction / heterotrimeric G-protein binding / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / ADP signalling through P2Y purinoceptor 1 / negative regulation of calcium ion-dependent exocytosis / G alpha (i) signalling events / positive regulation of blood pressure / Surfactant metabolism / positive regulation of potassium ion transport / dopaminergic synapse / thermoception / thioesterase binding / fear response / negative regulation of insulin secretion involved in cellular response to glucose stimulus / adenylate cyclase-inhibiting adrenergic receptor signaling pathway / norepinephrine binding / positive regulation of membrane protein ectodomain proteolysis / Adrenoceptors / intestinal absorption / response to alcohol / ADP signalling through P2Y purinoceptor 12 / Adrenaline,noradrenaline inhibits insulin secretion / Extra-nuclear estrogen signaling / photoreceptor outer segment membrane / adrenergic receptor signaling pathway / spectrin binding / G alpha (i) signalling events / positive regulation of wound healing / response to morphine / alkylglycerophosphoethanolamine phosphodiesterase activity / negative regulation of calcium ion transport / phototransduction, visible light / photoreceptor outer segment / regulation of vasoconstriction / negative regulation of lipid catabolic process / viral release from host cell by cytolysis / positive regulation of epidermal growth factor receptor signaling pathway / Rho protein signal transduction / cellular response to hormone stimulus / adenylate cyclase-activating adrenergic receptor signaling pathway / positive regulation of protein localization to cell cortex / cardiac muscle cell apoptotic process / peptidoglycan catabolic process / photoreceptor inner segment / positive regulation of neuron differentiation / presynaptic active zone membrane / G protein-coupled serotonin receptor binding / presynaptic modulation of chemical synaptic transmission / axon terminus / cellular response to forskolin / positive regulation of cytokine production / regulation of mitotic spindle organization / female pregnancy / platelet activation / negative regulation of insulin secretion / G protein-coupled receptor binding / postsynaptic density membrane / vasodilation / GABA-ergic synapse / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / epidermal growth factor receptor signaling pathway Similarity search - Function | |||||||||
| Biological species |  Spodoptera (butterflies/moths) / Spodoptera (butterflies/moths) /    Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Yuan D / Liu Z | |||||||||
 Citation Citation |  Journal: Nat Chem Biol / Year: 2020 Journal: Nat Chem Biol / Year: 2020Title: Activation of the α adrenoceptor by the sedative sympatholytic dexmedetomidine. Authors: Daopeng Yuan / Zhongmin Liu / Jonas Kaindl / Shoji Maeda / Jiawei Zhao / Xiaoou Sun / Jun Xu / Peter Gmeiner / Hong-Wei Wang / Brian K Kobilka /    Abstract: The α adrenergic receptors (αARs) are G protein-coupled receptors (GPCRs) that respond to adrenaline and noradrenaline and couple to the Gi/o family of G proteins. αARs play important roles in ...The α adrenergic receptors (αARs) are G protein-coupled receptors (GPCRs) that respond to adrenaline and noradrenaline and couple to the Gi/o family of G proteins. αARs play important roles in regulating the sympathetic nervous system. Dexmedetomidine is a highly selective αAR agonist used in post-operative patients as an anxiety-reducing, sedative medicine that decreases the requirement for opioids. As is typical for selective αAR agonists, dexmedetomidine consists of an imidazole ring and a substituted benzene moiety lacking polar groups, which is in contrast to βAR-selective agonists, which share an ethanolamine group and an aromatic system with polar, hydrogen-bonding substituents. To better understand the structural basis for the selectivity and efficacy of adrenergic agonists, we determined the structure of the αAR in complex with dexmedetomidine and Go at a resolution of 2.9 Å by single-particle cryo-EM. The structure reveals the mechanism of αAR-selective activation and provides insights into Gi/o coupling specificity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9912.map.gz emd_9912.map.gz | 3.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9912-v30.xml emd-9912-v30.xml emd-9912.xml emd-9912.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9912.png emd_9912.png | 48.3 KB | ||
| Filedesc metadata |  emd-9912.cif.gz emd-9912.cif.gz | 7.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9912 http://ftp.pdbj.org/pub/emdb/structures/EMD-9912 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9912 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9912 | HTTPS FTP |
-Related structure data
| Related structure data |  6k42MC  9911C  6k41C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9912.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9912.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | alpha2BAR-Gi1 complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.401 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : alpha2BAR-Gi1 complex
| Entire | Name: alpha2BAR-Gi1 complex |
|---|---|
| Components |
|
-Supramolecule #1: alpha2BAR-Gi1 complex
| Supramolecule | Name: alpha2BAR-Gi1 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Spodoptera (butterflies/moths) Spodoptera (butterflies/moths) |
| Molecular weight | Theoretical: 150 KDa |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 40.415031 KDa |
| Recombinant expression | Organism:  Spodoptera (butterflies/moths) Spodoptera (butterflies/moths) |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVGGQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHESM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCA TDTKNVQFVF DAVTDVIIKN NLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.530992 KDa |
| Recombinant expression | Organism:  Spodoptera (butterflies/moths) Spodoptera (butterflies/moths) |
| Sequence | String: HHHHHHGSSG QSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKL IIWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC R FLDDNQIV ...String: HHHHHHGSSG QSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKL IIWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC R FLDDNQIV TSSGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KLWDVREGMC RQTFTGHESD IN AICFFPN GNAFATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAG HDNRVS CLGVTDDGMA VATGSWDSFL KIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Spodoptera (butterflies/moths) Spodoptera (butterflies/moths) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: Alpha-2A adrenergic receptor,Endolysin,Alpha-2B adrenergic recept...
| Macromolecule | Name: Alpha-2A adrenergic receptor,Endolysin,Alpha-2B adrenergic receptor,Alpha-2B adrenergic receptor type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO / EC number: lysozyme |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 58.156211 KDa |
| Recombinant expression | Organism:  Spodoptera (butterflies/moths) Spodoptera (butterflies/moths) |
| Sequence | String: DDDDAHHHHH HMGSLQPDAG NASWNGTEAP GGGARATPEN LYFQGNIFEM LRIDEGLRLK IYKDTEGYYT IGIGHLLTKS PSLNAAKSE LDKAIGRNTN GVITKDEAEK LFNQDVDAAV RGILRNAKLK PVYDSLDAVR RAALINMVFQ MGETGVAGFT N SLRMLQQK ...String: DDDDAHHHHH HMGSLQPDAG NASWNGTEAP GGGARATPEN LYFQGNIFEM LRIDEGLRLK IYKDTEGYYT IGIGHLLTKS PSLNAAKSE LDKAIGRNTN GVITKDEAEK LFNQDVDAAV RGILRNAKLK PVYDSLDAVR RAALINMVFQ MGETGVAGFT N SLRMLQQK RWDEAAVNLA KSRWYNQTPN RAKRVITTFR TGTWDAYYSV QATAAIAAAI TFLILFTIFG NALVILAVLT SR SLRAPQN LFLVSLAAAD ILVATLIIPF SLANELLGYW YFRRTWCEVY LALDVLFCTS SIVHLCAISL DRYWAVSRAL EYN SKRTPR RIKCIILTVW LIAAVISLPP LIYKGDQGPQ PRGRPQCKLN QEAWYILASS IGSFFAPCLI MILVYLRIYL IAKR SNRRG PRAKGGPGQG EQWWRRRAQL TREKRFTFVL AVVIGVFVLC WFPFFFSYSL GAICPKHCKV PHGLFQFFFW IGYCN SSLN PVIYTIFNQD FRRAFRRILC RPWTQTAW UniProtKB: Alpha-2A adrenergic receptor, Endolysin, Alpha-2B adrenergic receptor, Alpha-2B adrenergic receptor |
-Macromolecule #5: scFv
| Macromolecule | Name: scFv / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 32.898781 KDa |
| Recombinant expression | Organism:  Spodoptera (butterflies/moths) Spodoptera (butterflies/moths) |
| Sequence | String: MLLVNQSHQG FNKEHTSKMV SAIVLYVLLA AAAHSAFADV QLVESGGGLV QPGGSRKLSC SASGFAFSSF GMHWVRQAPE KGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSEDT AMYYCVRSIY YYGSSPFDFW GQGTTLTVSS G GGGSGGGG ...String: MLLVNQSHQG FNKEHTSKMV SAIVLYVLLA AAAHSAFADV QLVESGGGLV QPGGSRKLSC SASGFAFSSF GMHWVRQAPE KGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSEDT AMYYCVRSIY YYGSSPFDFW GQGTTLTVSS G GGGSGGGG SGGGGSDIVM TQATSSVPVT PGESVSISCR SSKSLLHSNG NTYLYWFLQR PGQSPQLLIY RMSNLASGVP DR FSGSGSG TAFTLTISRL EAEDVGVYYC MQHLEYPLTF GAGTKLELKG SLEVLFQGPA AAHHHHHHHH |
-Macromolecule #6: 4-[(1~{S})-1-(2,3-dimethylphenyl)ethyl]-1~{H}-imidazole
| Macromolecule | Name: 4-[(1~{S})-1-(2,3-dimethylphenyl)ethyl]-1~{H}-imidazole type: ligand / ID: 6 / Number of copies: 1 / Formula: CZX |
|---|---|
| Molecular weight | Theoretical: 200.28 Da |
| Chemical component information | 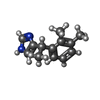 ChemComp-CZX: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 400 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















