+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24076 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
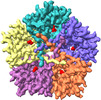
| Title | Cryo-EM structure of PfFNT-inhibitor complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PfFNT / Lactate transporter / PfFNT-MMV007839 complex / TRANSPORT PROTEIN-INHIBITOR complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationhigh-affinity secondary active nitrite transmembrane transporter activity / lactate transmembrane transport / nitrite transport / lactate:proton symporter activity / vacuolar membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.18 Å | |||||||||
 Authors Authors | Yu EW / Su C | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: EMBO Rep / Year: 2021 Journal: EMBO Rep / Year: 2021Title: Structural basis of transport and inhibition of the Plasmodium falciparum transporter PfFNT. Authors: Meinan Lyu / Chih-Chia Su / James W Kazura / Edward W Yu /  Abstract: The intra-erythrocyte stage of P. falciparum relies primarily on glycolysis to generate adenosine triphosphate (ATP) and the energy required to support growth and reproduction. Lactic acid, a ...The intra-erythrocyte stage of P. falciparum relies primarily on glycolysis to generate adenosine triphosphate (ATP) and the energy required to support growth and reproduction. Lactic acid, a metabolic byproduct of glycolysis, is potentially toxic as it lowers the pH inside the parasite. Plasmodium falciparum formate-nitrite transporter (PfFNT), a 34-kDa transmembrane protein, has been identified as a novel drug target as it exports lactate from inside the parasite to the surrounding parasitophorous vacuole within the erythrocyte cytosol. The structure and detailed molecular mechanism of this membrane protein are not yet available. Here we present structures of PfFNT in the absence and presence of the functional inhibitor MMV007839 at resolutions of 2.56 Å and 2.78 Å using single-particle cryo-electron microscopy. Genetic analysis and transport assay indicate that PfFNT is able to transfer lactate across the membrane. Combined, our data suggest a stepwise displacement mechanism for substrate transport. The PfFNT membrane protein is capable of picking up lactate ions from the parasite's cytosol, converting them to lactic acids and then exporting these acids into the extracellular space. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24076.map.gz emd_24076.map.gz | 3.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24076-v30.xml emd-24076-v30.xml emd-24076.xml emd-24076.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_24076.png emd_24076.png | 214.7 KB | ||
| Filedesc metadata |  emd-24076.cif.gz emd-24076.cif.gz | 5.6 KB | ||
| Others |  emd_24076_additional_1.map.gz emd_24076_additional_1.map.gz | 97.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24076 http://ftp.pdbj.org/pub/emdb/structures/EMD-24076 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24076 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24076 | HTTPS FTP |
-Related structure data
| Related structure data |  7mxyMC  6vqqC  6vqrC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24076.map.gz / Format: CCP4 / Size: 3.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24076.map.gz / Format: CCP4 / Size: 3.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: #1
| File | emd_24076_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PfFNT-inhibitor complex
| Entire | Name: PfFNT-inhibitor complex |
|---|---|
| Components |
|
-Supramolecule #1: PfFNT-inhibitor complex
| Supramolecule | Name: PfFNT-inhibitor complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Formate-nitrite transporter
| Macromolecule | Name: Formate-nitrite transporter / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 34.492281 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPPNNSKYVL DPVSIKSVCG GEESYIRCVE YGKKKAHYSN LNLLAKAILA GMFVGLCAHA SGIAGGLFYY HKLREIVGAS MSVFVYGFT FPIAFMCIIC TGSDLFTGNT LAVTMALYEK KVKLLDYLRV MTISLFGNYV GAVSFAFFVS YLSGAFTNVH A VEKNHFFQ ...String: MPPNNSKYVL DPVSIKSVCG GEESYIRCVE YGKKKAHYSN LNLLAKAILA GMFVGLCAHA SGIAGGLFYY HKLREIVGAS MSVFVYGFT FPIAFMCIIC TGSDLFTGNT LAVTMALYEK KVKLLDYLRV MTISLFGNYV GAVSFAFFVS YLSGAFTNVH A VEKNHFFQ FLNDIAEKKV HHTFVECVSL AVGCNIFVCL AVYFVLTLKD GAGYVFSVFF AVYAFAIAGY EHIIANIYTL NI ALMVNTK ITVYQAYIKN LLPTLLGNYI AGAIVLGLPL YFIYKEHYYN FERSKRDNND AQMKSLSIEL RN UniProtKB: Formate-nitrite transporter |
-Macromolecule #2: (Z)-4,4,5,5,5-pentakis(fluoranyl)-1-(4-methoxy-2-oxidanyl-phenyl)...
| Macromolecule | Name: (Z)-4,4,5,5,5-pentakis(fluoranyl)-1-(4-methoxy-2-oxidanyl-phenyl)-3-oxidanyl-pent-2-en-1-one type: ligand / ID: 2 / Number of copies: 5 / Formula: HV6 |
|---|---|
| Molecular weight | Theoretical: 312.189 Da |
| Chemical component information |  ChemComp-HV6: |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 2 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 38.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)