+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23020 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human mitochondrial LONP1 with endogenous substrate | |||||||||||||||
 Map data Map data | Substrate-bound human mitochondrial LONP1 | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | AAA+ / ATPase / protease / mitochondrial / LONP1 / LON / HYDROLASE | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationoxidation-dependent protein catabolic process / PH domain binding / mitochondrial protein catabolic process / G-quadruplex DNA binding / endopeptidase La / mitochondrial DNA metabolic process / mitochondrial genome maintenance / ATP-dependent peptidase activity / protein quality control for misfolded or incompletely synthesized proteins / mitochondrial nucleoid ...oxidation-dependent protein catabolic process / PH domain binding / mitochondrial protein catabolic process / G-quadruplex DNA binding / endopeptidase La / mitochondrial DNA metabolic process / mitochondrial genome maintenance / ATP-dependent peptidase activity / protein quality control for misfolded or incompletely synthesized proteins / mitochondrial nucleoid / regulation of peptidyl-tyrosine phosphorylation / insulin receptor substrate binding / chaperone-mediated protein complex assembly / DNA polymerase binding / negative regulation of insulin receptor signaling pathway / mitochondrion organization / proteolysis involved in protein catabolic process / Mitochondrial protein degradation / protein catabolic process / ADP binding / single-stranded DNA binding / cellular response to oxidative stress / sequence-specific DNA binding / single-stranded RNA binding / response to hypoxia / mitochondrial matrix / serine-type endopeptidase activity / ATP hydrolysis activity / mitochondrion / nucleoplasm / ATP binding / identical protein binding / membrane / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||||||||
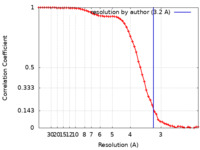
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||||||||
 Authors Authors | Shin M / Watson ER | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Structures of the human LONP1 protease reveal regulatory steps involved in protease activation. Authors: Mia Shin / Edmond R Watson / Albert S Song / Jeffrey T Mindrebo / Scott J Novick / Patrick R Griffin / R Luke Wiseman / Gabriel C Lander /  Abstract: The human mitochondrial AAA+ protein LONP1 is a critical quality control protease involved in regulating diverse aspects of mitochondrial biology including proteostasis, electron transport chain ...The human mitochondrial AAA+ protein LONP1 is a critical quality control protease involved in regulating diverse aspects of mitochondrial biology including proteostasis, electron transport chain activity, and mitochondrial transcription. As such, genetic or aging-associated imbalances in LONP1 activity are implicated in pathologic mitochondrial dysfunction associated with numerous human diseases. Despite this importance, the molecular basis for LONP1-dependent proteolytic activity remains poorly defined. Here, we solved cryo-electron microscopy structures of human LONP1 to reveal the underlying molecular mechanisms governing substrate proteolysis. We show that, like bacterial Lon, human LONP1 adopts both an open and closed spiral staircase orientation dictated by the presence of substrate and nucleotide. Unlike bacterial Lon, human LONP1 contains a second spiral staircase within its ATPase domain that engages substrate as it is translocated toward the proteolytic chamber. Intriguingly, and in contrast to its bacterial ortholog, substrate binding within the central ATPase channel of LONP1 alone is insufficient to induce the activated conformation of the protease domains. To successfully induce the active protease conformation in substrate-bound LONP1, substrate binding within the protease active site is necessary, which we demonstrate by adding bortezomib, a peptidomimetic active site inhibitor of LONP1. These results suggest LONP1 can decouple ATPase and protease activities depending on whether AAA+ or both AAA+ and protease domains bind substrate. Importantly, our structures provide a molecular framework to define the critical importance of LONP1 in regulating mitochondrial proteostasis in health and disease. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23020.map.gz emd_23020.map.gz | 14.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23020-v30.xml emd-23020-v30.xml emd-23020.xml emd-23020.xml | 25.2 KB 25.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_23020_fsc.xml emd_23020_fsc.xml | 7.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_23020.png emd_23020.png | 94.8 KB | ||
| Masks |  emd_23020_msk_1.map emd_23020_msk_1.map | 15.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-23020.cif.gz emd-23020.cif.gz | 7.5 KB | ||
| Others |  emd_23020_half_map_1.map.gz emd_23020_half_map_1.map.gz emd_23020_half_map_2.map.gz emd_23020_half_map_2.map.gz | 14.5 MB 14.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23020 http://ftp.pdbj.org/pub/emdb/structures/EMD-23020 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23020 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23020 | HTTPS FTP |
-Validation report
| Summary document |  emd_23020_validation.pdf.gz emd_23020_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_23020_full_validation.pdf.gz emd_23020_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_23020_validation.xml.gz emd_23020_validation.xml.gz | 11.5 KB | Display | |
| Data in CIF |  emd_23020_validation.cif.gz emd_23020_validation.cif.gz | 15.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23020 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23020 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23020 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23020 | HTTPS FTP |
-Related structure data
| Related structure data |  7ksmMC  7krzC  7kslC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23020.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23020.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Substrate-bound human mitochondrial LONP1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_23020_msk_1.map emd_23020_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
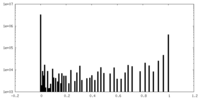
| Density Histograms |
-Half map: Half-map of substrate-bound human mitochondrial LONP1
| File | emd_23020_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map of substrate-bound human mitochondrial LONP1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
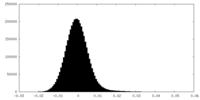
| Density Histograms |
-Half map: Half-map of substrate-bound human mitochondrial LONP1
| File | emd_23020_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map of substrate-bound human mitochondrial LONP1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
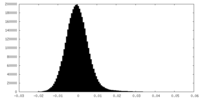
| Density Histograms |
- Sample components
Sample components
-Entire : Human mitochondrial LONP1 with endogenous substrate
| Entire | Name: Human mitochondrial LONP1 with endogenous substrate |
|---|---|
| Components |
|
-Supramolecule #1: Human mitochondrial LONP1 with endogenous substrate
| Supramolecule | Name: Human mitochondrial LONP1 with endogenous substrate / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 Details: Complexes consisting of homohexameric LONP1 protease from Homo sapiens bound to endogenous co-purified substrate |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organelle: Mitochondria / Location in cell: Matrix Homo sapiens (human) / Organelle: Mitochondria / Location in cell: Matrix |
| Molecular weight | Theoretical: 462 KDa |
-Macromolecule #1: Lon protease homolog, mitochondrial
| Macromolecule | Name: Lon protease homolog, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO / EC number: endopeptidase La |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 59.069742 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AIEEKFRERL KELVVPKHVM DVVDEELSKL GLLDNHSSEF NVTRNYLDWL TSIPWGKYSN ENLDLARAQA VLEEDHYGME DVKKRILEF IAVSQLRGST QGKILCFYGP PGVGKTSIAR SIARALNREY FRFSVGGMTD VAEIKGHRRT YVGAMPGKII Q CLKKTKTE ...String: AIEEKFRERL KELVVPKHVM DVVDEELSKL GLLDNHSSEF NVTRNYLDWL TSIPWGKYSN ENLDLARAQA VLEEDHYGME DVKKRILEF IAVSQLRGST QGKILCFYGP PGVGKTSIAR SIARALNREY FRFSVGGMTD VAEIKGHRRT YVGAMPGKII Q CLKKTKTE NPLILIDEVD KIGRGYQGDP SSALLELLDP EQNANFLDHY LDVPVDLSKV LFICTANVTD TIPEPLRDRM EM INVSGYV AQEKLAIAER YLVPQARALC GLDESKAKLS SDVLTLLIKQ YCRESGVRNL QKQVEKVLRK SAYKIVSGEA ESV EVTPEN LQDFVGKPVF TVERMYDVTP PGVVMGLAWT AMGGSTLFVE TSLRRPQDKD AKGDKDGSLE VTGQLGEVMK ESAR IAYTF ARAFLMQHAP ANDYLVTSHI HLHVPEGATP KDGPSAGCTI VTALLSLAMG RPVRQNLAMT GEVSLTGKIL PVGGI KEKT IAAKRAGVTC IVLPAENKKD FYDLAAFITE GLEVHFVEHY REIFDIAF UniProtKB: Lon protease homolog, mitochondrial |
-Macromolecule #2: Unidentified endogenous substrate
| Macromolecule | Name: Unidentified endogenous substrate / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.039273 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) |
-Macromolecule #3: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 4 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 4 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 2 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: Solutions were made fresh from concentrated and filtered using a 0.1 um syringe filter to avoid microbial contamination. Samples were mixed on ice and incubated for 5 minutes before vitrification. | ||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 7 sec. / Pretreatment - Atmosphere: OTHER Details: EM grids were plasma cleaned prior to sample application for 7 seconds using a Solarus plasma cleaner (Gatan, Inc.) with a 75% nitrogen, 25% oxygen atmosphere at 15W. | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: HOMEMADE PLUNGER Details: 4 uL of sample was applied per grid and manually blotted for 4 seconds followed by immediately plunge-freezing in liquid ethane cooled by liquid nitrogen.. | ||||||||||||||||||
| Details | This sample was monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Temperature | Min: 80.0 K / Max: 90.0 K |
| Alignment procedure | Coma free - Residual tilt: 0.14 mrad |
| Details | Coma-free alignment procedure from Herzik & Wu, Nature Methods (2017). Preliminary grid screening was performed manually prior to data collection. |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3710 pixel / Digitization - Dimensions - Height: 3838 pixel / Digitization - Frames/image: 0-113 / Number grids imaged: 1 / Number real images: 2912 / Average exposure time: 11.4 sec. / Average electron dose: 50.0 e/Å2 Details: Images were collected in counting mode at 10 frames per second. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 1.5 µm / Calibrated defocus min: 0.5 µm / Calibrated magnification: 43478 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 36000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Initial homology model was built using SWISS-MODEL and initial rigid body docking was done using UCSF Chimera. |
|---|---|
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 52 / Target criteria: Correlation coefficient |
| Output model |  PDB-7ksm: |
 Movie
Movie Controller
Controller













 Z
Z Y
Y X
X