[English] 日本語
 Yorodumi
Yorodumi- EMDB-21995: 1.8 Angstrom resolution structure of b-galactosidase with a 200 k... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21995 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 1.8 Angstrom resolution structure of b-galactosidase with a 200 kV cryoARM electron microscope | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | enzyme / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationalkali metal ion binding / lactose catabolic process / beta-galactosidase complex / beta-galactosidase / beta-galactosidase activity / carbohydrate binding / magnesium ion binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |   | |||||||||
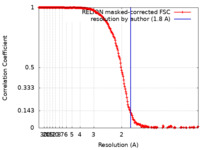
| Method | single particle reconstruction / cryo EM / Resolution: 1.8 Å | |||||||||
 Authors Authors | Merk A / Fukumura T | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: IUCrJ / Year: 2020 Journal: IUCrJ / Year: 2020Title: 1.8 Å resolution structure of β-galactosidase with a 200 kV CRYO ARM electron microscope. Authors: Alan Merk / Takuma Fukumura / Xing Zhu / Joseph E Darling / Reinhard Grisshammer / Jana Ognjenovic / Sriram Subramaniam /    Abstract: We report the determination of the structure of β-galactosidase at a resolution of ∼1.8 Å using data collected on a 200 kV CRYO ARM microscope equipped with a K3 direct electron detector. ...We report the determination of the structure of β-galactosidase at a resolution of ∼1.8 Å using data collected on a 200 kV CRYO ARM microscope equipped with a K3 direct electron detector. The data were collected in a single 24 h session by recording images from an array of 7 × 7 holes at each stage position using the automated data collection program . In addition to the expected features such as holes in the densities of aromatic residues, the map also shows density bumps corresponding to the locations of hydrogen atoms. The hydrogen densities are useful in assigning absolute orientations for residues such as glutamine or asparagine by removing the uncertainty in the fitting of the amide groups, and are likely to be especially relevant in the context of structure-guided drug design. These findings validate the use of electron microscopes operating at 200 kV for imaging protein complexes at atomic resolution using cryo-EM. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21995.map.gz emd_21995.map.gz | 1.8 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21995-v30.xml emd-21995-v30.xml emd-21995.xml emd-21995.xml | 17.5 KB 17.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21995_fsc.xml emd_21995_fsc.xml | 21.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_21995.png emd_21995.png | 216.6 KB | ||
| Masks |  emd_21995_msk_1.map emd_21995_msk_1.map | 824 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-21995.cif.gz emd-21995.cif.gz | 6.3 KB | ||
| Others |  emd_21995_half_map_1.map.gz emd_21995_half_map_1.map.gz emd_21995_half_map_2.map.gz emd_21995_half_map_2.map.gz | 667.3 MB 663.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21995 http://ftp.pdbj.org/pub/emdb/structures/EMD-21995 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21995 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21995 | HTTPS FTP |
-Related structure data
| Related structure data |  6x1qMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10446 (Title: 1.8 Å resolution structure of β-galactosidase with a 200 kV CRYO ARM electron microscope EMPIAR-10446 (Title: 1.8 Å resolution structure of β-galactosidase with a 200 kV CRYO ARM electron microscopeData size: 1.1 TB Data #1: uncorrected multi-frame micrographs of beta-galactosidase [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21995.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21995.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.26 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
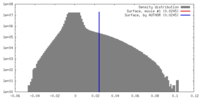
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_21995_msk_1.map emd_21995_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
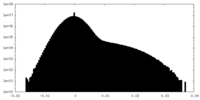
| Density Histograms |
-Half map: #1
| File | emd_21995_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
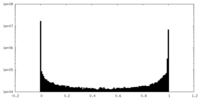
| Density Histograms |
-Half map: #2
| File | emd_21995_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
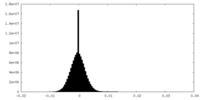
| Density Histograms |
- Sample components
Sample components
-Entire : beta-galactosidase
| Entire | Name: beta-galactosidase |
|---|---|
| Components |
|
-Supramolecule #1: beta-galactosidase
| Supramolecule | Name: beta-galactosidase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 465 KDa |
-Macromolecule #1: Beta-galactosidase
| Macromolecule | Name: Beta-galactosidase / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: beta-galactosidase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 116.181031 KDa |
| Sequence | String: MITDSLAVVL QRRDWENPGV TQLNRLAAHP PFASWRNSEE ARTDRPSQQL RSLNGEWRFA WFPAPEAVPE SWLECDLPEA DTVVVPSNW QMHGYDAPIY TNVTYPITVN PPFVPTENPT GCYSLTFNVD ESWLQEGQTR IIFDGVNSAF HLWCNGRWVG Y GQDSRLPS ...String: MITDSLAVVL QRRDWENPGV TQLNRLAAHP PFASWRNSEE ARTDRPSQQL RSLNGEWRFA WFPAPEAVPE SWLECDLPEA DTVVVPSNW QMHGYDAPIY TNVTYPITVN PPFVPTENPT GCYSLTFNVD ESWLQEGQTR IIFDGVNSAF HLWCNGRWVG Y GQDSRLPS EFDLSAFLRA GENRLAVMVL RWSDGSYLED QDMWRMSGIF RDVSLLHKPT TQISDFHVAT RFNDDFSRAV LE AEVQMCG ELRDYLRVTV SLWQGETQVA SGTAPFGGEI IDERGGYADR VTLRLNVENP KLWSAEIPNL YRAVVELHTA DGT LIEAEA CDVGFRVVRI ENGLLLLNGK PLLIRGVNRH EHHPLHGQVM DEQTMVQDIL LMKQNNFNAV RCSHYPNHPL WYTL CDRYG LYVVDEANIE THGMVPMNRL TDDPRWLPAM SERVTRMVQR DRNHPSVIIW SLGNESGHGA NHDALYRWIK SVDPS RPVQ YEGGGADTTA TDIICPMYAR VDEDQPFPAV PKWSIKKWLS LPGETRPLIL CEYAHAMGNS LGGFAKYWQA FRQYPR LQG GFVWDWVDQS LIKYDENGNP WSAYGGDFGD TPNDRQFCMN GLVFADRTPH PALTEAKHQQ QFFQFRLSGQ TIEVTSE YL FRHSDNELLH WMVALDGKPL ASGEVPLDVA PQGKQLIELP ELPQPESAGQ LWLTVRVVQP NATAWSEAGH ISAWQQWR L AENLSVTLPA ASHAIPHLTT SEMDFCIELG NKRWQFNRQS GFLSQMWIGD KKQLLTPLRD QFTRAPLDND IGVSEATRI DPNAWVERWK AAGHYQAEAA LLQCTADTLA DAVLITTAHA WQHQGKTLFI SRKTYRIDGS GQMAITVDVV VASDTPHPAR IGLNCQLAQ VAERVNWLGL GPQENYPDRL TAACFDRWDL PLSDMYTPYV FPSENGLRCG TRELNYGPHQ WRGDFQFNIS R YSQQQLME TSHRHLLHAE EGTWLNIDGF HMGIGGDDSW SPSVSAEFQL SAGRYHYQLV WCQ UniProtKB: Beta-galactosidase |
-Macromolecule #2: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 2 / Number of copies: 8 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #3: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 3 / Number of copies: 8 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 3618 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4.5 mg/mL |
|---|---|
| Buffer | pH: 8 / Details: 25mM Tris pH 8.0, 50mM NaCl, 0.5mM TCEP, 2mM MgCl |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 10 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 291 K / Instrument: LEICA EM GP |
| Details | Purified by size exclusion chromatography |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 200 |
|---|---|
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Slit width: 30 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 4949 / Average exposure time: 1.0 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: -0.9500000000000001 µm / Nominal defocus min: -0.8 µm |
| Sample stage | Specimen holder model: JEOL CRYOSPECPORTER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)