+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20668 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of the HCoV-229E spike glycoprotein | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CoV coronavirus 229E / spike glycoprotein / APN / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationhost cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / endocytosis involved in viral entry into host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion membrane / membrane Similarity search - Function | |||||||||
| Biological species |  Human coronavirus 229E Human coronavirus 229E | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Li Z / Benlekbir S / Rubinstein JL / Rini JM | |||||||||
| Funding support |  Canada, 1 items Canada, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2019 Journal: Elife / Year: 2019Title: The human coronavirus HCoV-229E S-protein structure and receptor binding. Authors: Zhijie Li / Aidan Ca Tomlinson / Alan Hm Wong / Dongxia Zhou / Marc Desforges / Pierre J Talbot / Samir Benlekbir / John L Rubinstein / James M Rini /  Abstract: The coronavirus S-protein mediates receptor binding and fusion of the viral and host cell membranes. In HCoV-229E, its receptor binding domain (RBD) shows extensive sequence variation but how S- ...The coronavirus S-protein mediates receptor binding and fusion of the viral and host cell membranes. In HCoV-229E, its receptor binding domain (RBD) shows extensive sequence variation but how S-protein function is maintained is not understood. Reported are the X-ray crystal structures of Class III-V RBDs in complex with human aminopeptidase N (hAPN), as well as the electron cryomicroscopy structure of the 229E S-protein. The structures show that common core interactions define the specificity for hAPN and that the peripheral RBD sequence variation is accommodated by loop plasticity. The results provide insight into immune evasion and the cross-species transmission of 229E and related coronaviruses. We also find that the 229E S-protein can expose a portion of its helical core to solvent. This is undoubtedly facilitated by hydrophilic subunit interfaces that we show are conserved among coronaviruses. These interfaces likely play a role in the S-protein conformational changes associated with membrane fusion. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20668.map.gz emd_20668.map.gz | 42.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20668-v30.xml emd-20668-v30.xml emd-20668.xml emd-20668.xml | 20.2 KB 20.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20668.png emd_20668.png | 95.1 KB | ||
| Masks |  emd_20668_msk_1.map emd_20668_msk_1.map | 83.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-20668.cif.gz emd-20668.cif.gz | 7 KB | ||
| Others |  emd_20668_additional.map.gz emd_20668_additional.map.gz emd_20668_half_map_1.map.gz emd_20668_half_map_1.map.gz emd_20668_half_map_2.map.gz emd_20668_half_map_2.map.gz | 79 MB 77.8 MB 77.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20668 http://ftp.pdbj.org/pub/emdb/structures/EMD-20668 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20668 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20668 | HTTPS FTP |
-Related structure data
| Related structure data |  6u7hMC  6u7eC  6u7fC  6u7gC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20668.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20668.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
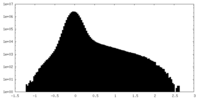
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_20668_msk_1.map emd_20668_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Sharpened map
| File | emd_20668_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_20668_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_20668_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human coronavirus 229E spike glycoprotein
| Entire | Name: Human coronavirus 229E spike glycoprotein |
|---|---|
| Components |
|
-Supramolecule #1: Human coronavirus 229E spike glycoprotein
| Supramolecule | Name: Human coronavirus 229E spike glycoprotein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: Ectodomain generated by recombinant expression in HEK293 Freestyle cells |
|---|---|
| Source (natural) | Organism:  Human coronavirus 229E Human coronavirus 229E |
| Molecular weight | Theoretical: 380 KDa |
-Macromolecule #1: spike glycoprotein
| Macromolecule | Name: spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human coronavirus 229E / Strain: VR-740 Human coronavirus 229E / Strain: VR-740 |
| Molecular weight | Theoretical: 127.231117 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVLLVAYAL LHIAGCQTTN GLNTSYSVCN GCVGYSENVF AVESGGYIPS DFAFNNWFLL TNTSSVVDGV VRSFQPLLLN CLWSVSGLR FTTGFVYFNG TGRGDCKGFS SDVLSDVIRY NLNFEENLRR GTILFKTSYG VVVFYCTNNT LVSGDAHIPF G TVLGNFYC ...String: MFVLLVAYAL LHIAGCQTTN GLNTSYSVCN GCVGYSENVF AVESGGYIPS DFAFNNWFLL TNTSSVVDGV VRSFQPLLLN CLWSVSGLR FTTGFVYFNG TGRGDCKGFS SDVLSDVIRY NLNFEENLRR GTILFKTSYG VVVFYCTNNT LVSGDAHIPF G TVLGNFYC FVNTTIGTET TSAFVGALPK TVREFVISRT GHFYINGYRY FTLGNVEAVN FNVTTAETTD FFTVALASYA DV LVNVSQT SIANIIYCNS VINRLRCDQL SFYVPDGFYS TSPIQSVELP VSIVSLPVYH KHMFIVLYVD FKPQSGGGKC FNC YPAGVN ITLANFNETK GPLCVDTSHF TTKYVAVYAN VGRWSASINT GNCPFSFGKV NNFVKFGSVC FSLKDIPGGC AMPI VANWA YSKYYTIGTL YVSWSDGDGI TGVPQPVEGV SSFMNVTLDK CTKYNIYDVS GVGVIRVSND TFLNGITYTS TSGNL LGFK DVTKGTIYSI TPCNPPDQLV VYQQAVVGAM LSENFTSYGF SNVVELPKFF YASNGTYNCT DAVLTYSSFG VCADGS IIA VQPRNVSYDS VSAIVTANLS IPSNWTISVQ VEYLQITSTP IVVDCSTYVC NGNVRCVELL KQYTSACKTI EDALRNS AR LESADVSEML TFDKKAFTLA NVSSFGDYNL SSVIPSLPTS GSRVAGRSAI EDILFSKIVT SGLGTVDADY KNCTKGLS I ADLACAQYYN GIMVLPGVAD AERMAMYTGS LIGGIALGGL TSAVSIPFSL AIQARLNYVA LQTDVLQENQ KILAASFNK AMTNIVDAFT GVNDAITQTS QALQTVATAL NKIQDVVNQQ GNSLNHLTSQ LRQNFQAISS SIQAIYDRLD PPQADQQVDR LITGRLAAL NVFVSHTLTK YTEVRASRQL AQQKVNECVK SQSKRYGFCG NGTHIFSIVN AAPEGLVFLH TVLLPTQYKD V EAWSGLCV DGTNGYVLRQ PNLALYKEGN YYRITSRIMF EPRIPTMADF VQIENCNVTF VNISRSELQT IVPEYIDVNK TL QELSYKL PNYTVPDLVV EQYNQTILNL TSEISTLENK SAELNYTVQK LQTLIDNINS TLVDLKWLNR VETYIKSGGY IPE APRDGQ AYVRKDGEWV LLSTFLNSEN LYFQSGSHHH HHH |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 33 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Material: COPPER / Mesh: 400 / Support film - topology: HOLEY ARRAY / Support film - Film thickness: 3.5 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 120 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.04 kPa / Details: 15 micro Ampere | |||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: GATAN CRYOPLUNGE 3 / Details: Blot for 13 seconds before plunging. | |||||||||
| Details | This sample was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average exposure time: 60.0 sec. / Average electron dose: 42.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE Details: A initial 3D map was generated by ab-initio reconstruction in cryoSPARC v2. |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 3.1 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 2.90) / Number images used: 71350 |
| Initial angle assignment | Type: OTHER |
| Final angle assignment | Type: OTHER |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | Manual model building was done in COOT and ChimeraX/ISOLDE. A homology model based on PDB:5SZS was used as a reference during model building. |
| Refinement | Space: REAL / Protocol: OTHER |
| Output model |  PDB-6u7h: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)