[English] 日本語
 Yorodumi
Yorodumi- EMDB-19587: CryoEM structure of M. smegmatis GMP reductase in complex with GM... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of M. smegmatis GMP reductase in complex with GMP and ATP at pH 6.6, compressed conformation. | |||||||||
 Map data Map data | Loc-Scale filtered | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GMP reductase / GuaB1 / CBS domain / Mycobacterium smegmatis / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationGMP reductase / GMP reductase activity / IMP salvage / IMP dehydrogenase activity / purine ribonucleoside salvage / cytosol Similarity search - Function | |||||||||
| Biological species |  Mycolicibacterium smegmatis (bacteria) Mycolicibacterium smegmatis (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.48 Å | |||||||||
 Authors Authors | Dolezal M / Kouba T / Pichova I | |||||||||
| Funding support | European Union, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2026 Journal: Nat Commun / Year: 2026Title: Structural basis for allosteric regulation of mycobacterial guanosine 5´-monophosphate reductase by ATP and GTP. Authors: Michal Doležal / Zdeněk Knejzlík / Tomáš Kouba / Anatolij Filimoněnko / Hana Šváchová / Matteo Dedola / Martin Klíma / Iva Pichová /  Abstract: Guanosine 5'-monophosphate reductase (GMPR) is a crucial enzyme in the purine salvage pathway that catalyses the NADPH-dependent conversion of GMP to IMP, thereby contributing to purine nucleotide ...Guanosine 5'-monophosphate reductase (GMPR) is a crucial enzyme in the purine salvage pathway that catalyses the NADPH-dependent conversion of GMP to IMP, thereby contributing to purine nucleotide homeostasis. Mycobacterium smegmatis GMPR (MsmGMPR) contains a regulatory cystathionine β-synthase (CBS) domain, which mediates allosteric modulation by ATP and GTP. However, MsmGMPR exhibits an atypical tertiary structure that is incompatible with the acknowledged regulatory mechanisms of IMPDH/GMPR family enzymes. Here, we combine X-ray crystallography, cryogenic electron microscopy, and biochemical binding assays to elucidate the molecular basis of MsmGMPR regulation by ATP and GTP. We show that ATP stabilises a compressed conformation that inhibits the enzyme by restricting access to the active site and preventing NADPH binding. In contrast, GTP counteracts ATP binding, promoting an active conformation that enables catalysis. Our results provide insight into how MsmGMPR senses and responds to the purine nucleotide balance, revealing a distinct utilisation of the CBS domain compared with its typical role in IMPDH/GMPR enzymes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19587.map.gz emd_19587.map.gz | 159.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19587-v30.xml emd-19587-v30.xml emd-19587.xml emd-19587.xml | 23.9 KB 23.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19587_fsc.xml emd_19587_fsc.xml | 15.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_19587.png emd_19587.png | 183.9 KB | ||
| Masks |  emd_19587_msk_1.map emd_19587_msk_1.map | 325 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-19587.cif.gz emd-19587.cif.gz | 6.9 KB | ||
| Others |  emd_19587_additional_1.map.gz emd_19587_additional_1.map.gz emd_19587_half_map_1.map.gz emd_19587_half_map_1.map.gz emd_19587_half_map_2.map.gz emd_19587_half_map_2.map.gz | 18 MB 257.3 MB 257.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19587 http://ftp.pdbj.org/pub/emdb/structures/EMD-19587 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19587 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19587 | HTTPS FTP |
-Related structure data
| Related structure data |  8ry5MC  8ry0C  8ry1C  8ry3C  8ry4C  8ry6C  8ry7C  8ry8C  8ry9C  8ryaC  8rybC  9hfzC  9hg0C  9hg1C  9hg2C  9hg3C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19587.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19587.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Loc-Scale filtered | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8336 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19587_msk_1.map emd_19587_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: RELION post-processed and masked
| File | emd_19587_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RELION post-processed and masked | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_19587_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_19587_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Msm GMPR with GMP and ATP at pH 6.6
| Entire | Name: Msm GMPR with GMP and ATP at pH 6.6 |
|---|---|
| Components |
|
-Supramolecule #1: Msm GMPR with GMP and ATP at pH 6.6
| Supramolecule | Name: Msm GMPR with GMP and ATP at pH 6.6 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 |
-Macromolecule #1: GMP reductase
| Macromolecule | Name: GMP reductase / type: protein_or_peptide / ID: 1 / Details: Guanosine 5'-monophosphate reductase / Number of copies: 8 / Enantiomer: LEVO / EC number: GMP reductase |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 |
| Molecular weight | Theoretical: 51.782434 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVRFLDGHTP AYDLTYNDVF VVPGRSDVAS RFDVDLSTVD GSGTTIPVVV ANMTAVAGRR MAETVARRGG IVVLPQDLPI TAVSETVDF VKSRDLVVDT PVTLSPEDSV SDANALLHKR AHGAAVVVFE GRPIGLVTEA NCAGVDRFAR VRDIALSDFV T APVGTDPR ...String: MVRFLDGHTP AYDLTYNDVF VVPGRSDVAS RFDVDLSTVD GSGTTIPVVV ANMTAVAGRR MAETVARRGG IVVLPQDLPI TAVSETVDF VKSRDLVVDT PVTLSPEDSV SDANALLHKR AHGAAVVVFE GRPIGLVTEA NCAGVDRFAR VRDIALSDFV T APVGTDPR EVFDLLEHAP IDVAVMTAPD GTLAGVLTRT GAIRAGIYTP AVDAKGRLRI AAAVGINGDV GAKAQALAEA GA DLLVIDT AHGHQAKMLD AIKAVASLDL GLPLVAGNVV SAEGTRDLIE AGASIVKVGV GPGAMCTTRM MTGVGRPQFS AVV ECAAAA RQLGGHVWAD GGVRHPRDVA LALAAGASNV MIGSWFAGTY ESPGDLLFDR DDRPYKESYG MASKRAVAAR TAGD SSFDR ARKGLFEEGI STSRMSLDPA RGGVEDLLDH ITSGVRSTCT YVGAANLPEL HEKVVLGVQS AAGFAEGHPL PAGWT AAAK EDLEHHHHHH HH UniProtKB: GMP reductase |
-Macromolecule #2: GUANOSINE-5'-MONOPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-MONOPHOSPHATE / type: ligand / ID: 2 / Number of copies: 8 / Formula: 5GP |
|---|---|
| Molecular weight | Theoretical: 363.221 Da |
| Chemical component information | 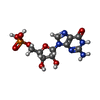 ChemComp-5GP: |
-Macromolecule #3: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 8 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 214 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.6 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE | ||||||||||||
| Details | 20 mg/ml Msm GMPR with 2 mM GMP and 5 mM ATP in the storage buffer (50 mM Tris, pH 8.0, 2.5 mM TCEP) was diluted with the cryoEM buffer (50 mM HEPES, pH 6.6, 100 mM KCl, 2 mM MgCl2). |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average exposure time: 2.0 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 30.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.9 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | The initial structure was obtained by fitting an appropriate model into the cryo-EM map using MolRep and ChimeraX. The structure was then refined by iterative manual rebuilding in Coot and Isolde, and automatic refinement in phenix.real_space_refine. |
|---|---|
| Refinement | Space: REAL |
| Output model |  PDB-8ry5: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















































