[English] 日本語
 Yorodumi
Yorodumi- EMDB-17558: Cryo-EM structure of cortactin stabilized Arp2/3-complex nucleate... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of cortactin stabilized Arp2/3-complex nucleated actin branches | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / CONTRACTILE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcytoskeletal calyx / tubulobulbar complex / meiotic chromosome movement towards spindle pole / cytosolic transport / growth cone leading edge / lamellipodium organization / Advanced glycosylation endproduct receptor signaling / RHOF GTPase cycle / spindle localization / meiotic cytokinesis ...cytoskeletal calyx / tubulobulbar complex / meiotic chromosome movement towards spindle pole / cytosolic transport / growth cone leading edge / lamellipodium organization / Advanced glycosylation endproduct receptor signaling / RHOF GTPase cycle / spindle localization / meiotic cytokinesis / site of polarized growth / : / actin polymerization-dependent cell motility / Regulation of actin dynamics for phagocytic cup formation / EPHB-mediated forward signaling / Adherens junctions interactions / VEGFA-VEGFR2 Pathway / Cell-extracellular matrix interactions / RHO GTPases Activate WASPs and WAVEs / MAP2K and MAPK activation / Formation of the canonical BAF (cBAF) complex / Formation of the polybromo-BAF (pBAF) complex / Formation of the embryonic stem cell BAF (esBAF) complex / Formation of the non-canonical BAF (ncBAF) complex / UCH proteinases / muscle cell projection membrane / negative regulation of filopodium assembly / Gap junction degradation / Formation of annular gap junctions / RHOF GTPase cycle / COPI-independent Golgi-to-ER retrograde traffic / Clathrin-mediated endocytosis / Regulation of CDH1 Function / Formation of the dystrophin-glycoprotein complex (DGC) / asymmetric cell division / Arp2/3 protein complex / HSP90 chaperone cycle for steroid hormone receptors (SHR) in the presence of ligand / actin nucleation / Arp2/3 complex-mediated actin nucleation / COPI-mediated anterograde transport / Arp2/3 complex binding / mitotic spindle midzone / F-actin capping protein complex / WASH complex / regulation of cell projection assembly / actin cap / sperm head / modification of postsynaptic actin cytoskeleton / regulation of mitophagy / profilin binding / Factors involved in megakaryocyte development and platelet production / modification of postsynaptic structure / cellular response to cytochalasin B / regulation of actin filament polymerization / positive regulation of smooth muscle contraction / substrate-dependent cell migration, cell extension / regulation of transepithelial transport / focal adhesion assembly / morphogenesis of a polarized epithelium / structural constituent of postsynaptic actin cytoskeleton / cell junction assembly / positive regulation of chemotaxis / protein localization to adherens junction / MHC class II antigen presentation / dense body / barbed-end actin filament capping / muscle cell development / Tat protein binding / regulation of lamellipodium assembly / actin polymerization or depolymerization / postsynaptic actin cytoskeleton / proline-rich region binding / podosome / regulation of cell morphogenesis / adherens junction assembly / apical protein localization / cell projection organization / dendritic spine maintenance / RHO GTPases activate IQGAPs / RHO GTPases Activate Formins / regulation of axon extension / tight junction / lamellipodium assembly / apical junction complex / cortical actin cytoskeleton / negative regulation of microtubule polymerization / regulation of norepinephrine uptake / transporter regulator activity / NuA4 histone acetyltransferase complex / filamentous actin / positive regulation of actin filament polymerization / establishment or maintenance of cell polarity / cortical cytoskeleton / nitric-oxide synthase binding / brush border / intercalated disc / cilium assembly / sperm head-tail coupling apparatus / RHO GTPases Activate WASPs and WAVEs / regulation of synaptic vesicle endocytosis Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /    Amanita phalloides (death cap) Amanita phalloides (death cap) | |||||||||
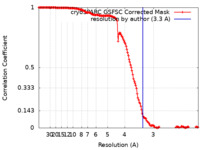
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Liu T / Moores CA | |||||||||
| Funding support | European Union, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Cortactin stabilizes actin branches by bridging activated Arp2/3 to its nucleated actin filament. Authors: Tianyang Liu / Luyan Cao / Miroslav Mladenov / Antoine Jegou / Michael Way / Carolyn A Moores /   Abstract: Regulation of the assembly and turnover of branched actin filament networks nucleated by the Arp2/3 complex is essential during many cellular processes, including cell migration and membrane ...Regulation of the assembly and turnover of branched actin filament networks nucleated by the Arp2/3 complex is essential during many cellular processes, including cell migration and membrane trafficking. Cortactin is important for actin branch stabilization, but the mechanism by which this occurs is unclear. Given this, we determined the structure of vertebrate cortactin-stabilized Arp2/3 actin branches using cryogenic electron microscopy. We find that cortactin interacts with the new daughter filament nucleated by the Arp2/3 complex at the branch site, rather than the initial mother actin filament. Cortactin preferentially binds activated Arp3. It also stabilizes the F-actin-like interface of activated Arp3 with the first actin subunit of the new filament, and its central repeats extend along successive daughter-filament subunits. The preference of cortactin for activated Arp3 explains its retention at the actin branch and accounts for its synergy with other nucleation-promoting factors in regulating branched actin network dynamics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17558.map.gz emd_17558.map.gz | 163.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17558-v30.xml emd-17558-v30.xml emd-17558.xml emd-17558.xml | 34.6 KB 34.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17558_fsc.xml emd_17558_fsc.xml | 14.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_17558.png emd_17558.png | 72 KB | ||
| Filedesc metadata |  emd-17558.cif.gz emd-17558.cif.gz | 9.3 KB | ||
| Others |  emd_17558_half_map_1.map.gz emd_17558_half_map_1.map.gz emd_17558_half_map_2.map.gz emd_17558_half_map_2.map.gz | 301.6 MB 301.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17558 http://ftp.pdbj.org/pub/emdb/structures/EMD-17558 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17558 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17558 | HTTPS FTP |
-Related structure data
| Related structure data |  8p94MC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17558.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17558.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.067 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_17558_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
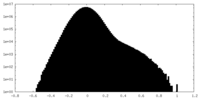
| Density Histograms |
-Half map: #1
| File | emd_17558_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Cortactin stabilizes Arp2/3-complex nucleated actin branches with...
+Supramolecule #1: Cortactin stabilizes Arp2/3-complex nucleated actin branches with...
+Supramolecule #2: Human Arp2/3 C1BC5L complex
+Supramolecule #3: Porcine actin, cytoplasmic 1
+Supramolecule #4: Mouse cortactin
+Supramolecule #5: Mouse capping protein
+Supramolecule #6: Phalloidin
+Macromolecule #1: Actin-related protein 3
+Macromolecule #2: Actin-related protein 2
+Macromolecule #3: Actin-related protein 2/3 complex subunit 1B
+Macromolecule #4: Actin-related protein 2/3 complex subunit 2
+Macromolecule #5: Actin-related protein 2/3 complex subunit 3
+Macromolecule #6: Actin-related protein 2/3 complex subunit 4
+Macromolecule #7: Actin-related protein 2/3 complex subunit 5-like protein
+Macromolecule #8: Actin, cytoplasmic 1
+Macromolecule #9: Src substrate cortactin
+Macromolecule #10: F-actin-capping protein subunit alpha-1
+Macromolecule #11: Isoform 2 of F-actin-capping protein subunit beta
+Macromolecule #12: Phalloidin
+Macromolecule #13: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #14: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 20 mM HEPES pH 7.5, 50mM KCl, 1mM EGTA, 1mM MgCl2, 0.2 mM ATP and 1 mM DTT |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 98 % / Chamber temperature: 295.15 K / Instrument: LEICA EM GP / Details: Back blotting. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 49.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 81000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Refinement | Protocol: OTHER |
| Output model |  PDB-8p94: |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)



































 unidentified baculovirus
unidentified baculovirus