[English] 日本語
 Yorodumi
Yorodumi- EMDB-17320: In situ subtomogram average of the Prototype Foamy Virus capsid, ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | In situ subtomogram average of the Prototype Foamy Virus capsid, p68 Gag | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | capsid / Gag / foamy virus / p68 / VIRAL PROTEIN | |||||||||
| Biological species |  Eastern chimpanzee simian foamy virus Eastern chimpanzee simian foamy virus | |||||||||
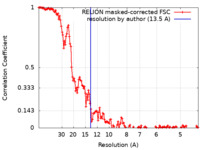
| Method | subtomogram averaging / cryo EM / Resolution: 13.5 Å | |||||||||
 Authors Authors | Calcraft T / Nans A / Rosenthal PB | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2024 Journal: Cell / Year: 2024Title: Integrated cryoEM structure of a spumaretrovirus reveals cross-kingdom evolutionary relationships and the molecular basis for assembly and virus entry. Authors: Thomas Calcraft / Nicole Stanke-Scheffler / Andrea Nans / Dirk Lindemann / Ian A Taylor / Peter B Rosenthal /   Abstract: Foamy viruses (FVs) are an ancient lineage of retroviruses, with an evolutionary history spanning over 450 million years. Vector systems based on Prototype Foamy Virus (PFV) are promising candidates ...Foamy viruses (FVs) are an ancient lineage of retroviruses, with an evolutionary history spanning over 450 million years. Vector systems based on Prototype Foamy Virus (PFV) are promising candidates for gene and oncolytic therapies. Structural studies of PFV contribute to the understanding of the mechanisms of FV replication, cell entry and infection, and retroviral evolution. Here we combine cryoEM and cryoET to determine high-resolution in situ structures of the PFV icosahedral capsid (CA) and envelope glycoprotein (Env), including its type III transmembrane anchor and membrane-proximal external region (MPER), and show how they are organized in an integrated structure of assembled PFV particles. The atomic models reveal an ancient retroviral capsid architecture and an unexpected relationship between Env and other class 1 fusion proteins of the Mononegavirales. Our results represent the de novo structure determination of an assembled retrovirus particle. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17320.map.gz emd_17320.map.gz | 228.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17320-v30.xml emd-17320-v30.xml emd-17320.xml emd-17320.xml | 18.1 KB 18.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17320_fsc.xml emd_17320_fsc.xml | 14.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_17320.png emd_17320.png | 71.1 KB | ||
| Masks |  emd_17320_msk_1.map emd_17320_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17320.cif.gz emd-17320.cif.gz | 5.6 KB | ||
| Others |  emd_17320_half_map_1.map.gz emd_17320_half_map_1.map.gz emd_17320_half_map_2.map.gz emd_17320_half_map_2.map.gz | 120 MB 120 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17320 http://ftp.pdbj.org/pub/emdb/structures/EMD-17320 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17320 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17320 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17320.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17320.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
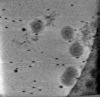
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.22 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17320_msk_1.map emd_17320_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
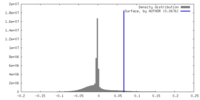
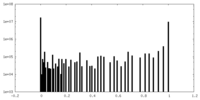
| Density Histograms |
-Half map: #1
| File | emd_17320_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_17320_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Eastern chimpanzee simian foamy virus
| Entire | Name:  Eastern chimpanzee simian foamy virus Eastern chimpanzee simian foamy virus |
|---|---|
| Components |
|
-Supramolecule #1: Eastern chimpanzee simian foamy virus
| Supramolecule | Name: Eastern chimpanzee simian foamy virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 2170195 / Sci species name: Eastern chimpanzee simian foamy virus / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Virus shell | Shell ID: 1 / T number (triangulation number): 13 |
-Macromolecule #1: Gag protein
| Macromolecule | Name: Gag protein / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Eastern chimpanzee simian foamy virus Eastern chimpanzee simian foamy virus |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MASGSNVEEY ELDVEALVVI LRDRNIPRNP LHGEVIGLRL TEGWWGQIER FQMVRLILQN DDNEPLQRPR YEVIQRAVNP HTMFMISGPL AELQLAFQDL DLPEGPLRFG PLANGHYVQG DPYSSSYRPV TMAETAQMTR DELEDVLNTQ SEIEIQMINL LELYEVETRA ...String: MASGSNVEEY ELDVEALVVI LRDRNIPRNP LHGEVIGLRL TEGWWGQIER FQMVRLILQN DDNEPLQRPR YEVIQRAVNP HTMFMISGPL AELQLAFQDL DLPEGPLRFG PLANGHYVQG DPYSSSYRPV TMAETAQMTR DELEDVLNTQ SEIEIQMINL LELYEVETRA LRRQLAERSS TGQGGISPGA PRSRPPVSSF SGLPSLPSIP GIHPRAPSPP RATSTPGNIP WSLGDDNPPS SSFPGPSQPR VSFHPGNPFV EEEGHRPRSQ SRERRREILP APVPSAPPMI QYIPVPPPPP IGTVIPIQHI RSVTGEPPRN PREIPIWLGR NAPAIDGVFP VTTPDLRCRI INAILGGNIG LSLTPGDCLT WDSAVATLFI RTHGTFPMHQ LGNVIKGIVD QEGVATAYTL GMMLSGQNYQ LVSGIIRGYL PGQAVVTALQ QRLDQEIDDQ TRAETFIQHL NAVYEILGLN ARGQSIRASV TPQPRPSRGR GRGQNTSRPS QGPANSGRGR QRPASGQSNR GSSTQNQNQD NLNQGGYNLR PRTYQPQRYG GGRGRRWNDN TNNQESRPSD QGSQTPRPNQ AGSGVRGNQS QTPRPAAGRG GRGNHNRNQR SSGAGDSRAV N |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Average electron dose: 1.65 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.5 µm / Nominal defocus min: 2.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)