+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Photorhabdus luminescens TcdA1 prepore-to-pore intermediate, K567W K2008W mutant | |||||||||
 マップデータ マップデータ | The local filtered map that was used for model building | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Bacterial toxin / Tc toxin / TOXIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報 | |||||||||
| 生物種 |  Photorhabdus luminescens (バクテリア) Photorhabdus luminescens (バクテリア) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.2 Å | |||||||||
 データ登録者 データ登録者 | Nganga PN / Roderer D / Belyy A / Prumbaum D / Raunser S | |||||||||
| 資金援助 |  ドイツ, 1件 ドイツ, 1件
| |||||||||
 引用 引用 |  ジャーナル: Sci Adv / 年: 2025 ジャーナル: Sci Adv / 年: 2025タイトル: Multistate kinetics of the syringe-like injection mechanism of Tc toxins. 著者: Peter Njenga Ng'ang'a / Julian Folz / Svetlana Kucher / Daniel Roderer / Ying Xu / Oleg Sitsel / Alexander Belyy / Daniel Prumbaum / Ralf Kühnemuth / Tufa E Assafa / Min Dong / Claus A M ...著者: Peter Njenga Ng'ang'a / Julian Folz / Svetlana Kucher / Daniel Roderer / Ying Xu / Oleg Sitsel / Alexander Belyy / Daniel Prumbaum / Ralf Kühnemuth / Tufa E Assafa / Min Dong / Claus A M Seidel / Enrica Bordignon / Stefan Raunser /    要旨: Tc toxins are pore-forming virulence factors of many pathogenic bacteria. Following pH-induced conformational changes, they perforate the target membrane like a syringe to translocate toxic enzymes ...Tc toxins are pore-forming virulence factors of many pathogenic bacteria. Following pH-induced conformational changes, they perforate the target membrane like a syringe to translocate toxic enzymes into a cell. Although this complex transformation has been structurally well studied, the reaction pathway and the resulting temporal evolution have remained elusive. We used an integrated biophysical approach to monitor prepore-to-pore transition and found a reaction time of ~30 hours for a complete transition. We show two asynchronous general steps of the process, shell opening and channel ejection, with the overall reaction pathway being a slow multistep process involving three intermediates. Liposomes, an increasingly high pH, or receptors facilitate shell opening, which is directly correlated with an increased rate of the prepore-to-pore transition. Channel ejection is a near-instantaneous process which occurs with a transition time of <60 milliseconds. Understanding the mechanism of action of Tc toxins and unveiling modulators of the kinetics are key steps toward their application as biomedical devices or biopesticides. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_16792.map.gz emd_16792.map.gz | 19.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-16792-v30.xml emd-16792-v30.xml emd-16792.xml emd-16792.xml | 19.8 KB 19.8 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_16792.png emd_16792.png | 123 KB | ||
| マスクデータ |  emd_16792_msk_1.map emd_16792_msk_1.map | 669.9 MB |  マスクマップ マスクマップ | |
| Filedesc metadata |  emd-16792.cif.gz emd-16792.cif.gz | 8 KB | ||
| その他 |  emd_16792_half_map_1.map.gz emd_16792_half_map_1.map.gz emd_16792_half_map_2.map.gz emd_16792_half_map_2.map.gz | 541.1 MB 540.2 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16792 http://ftp.pdbj.org/pub/emdb/structures/EMD-16792 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16792 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16792 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_16792.map.gz / 形式: CCP4 / 大きさ: 669.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_16792.map.gz / 形式: CCP4 / 大きさ: 669.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | The local filtered map that was used for model building | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.88 Å | ||||||||||||||||||||||||||||||||||||
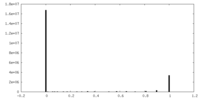
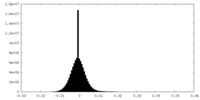
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-マスク #1
| ファイル |  emd_16792_msk_1.map emd_16792_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
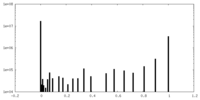
| 投影像・断面図 |
| ||||||||||||
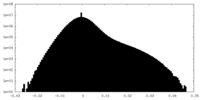
| 密度ヒストグラム |
-ハーフマップ: First half map
| ファイル | emd_16792_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | First half map | ||||||||||||
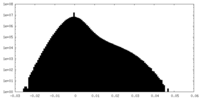
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: Second half map
| ファイル | emd_16792_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Second half map | ||||||||||||
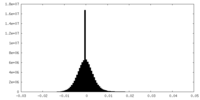
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : TcdA1
| 全体 | 名称: TcdA1 |
|---|---|
| 要素 |
|
-超分子 #1: TcdA1
| 超分子 | 名称: TcdA1 / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  Photorhabdus luminescens (バクテリア) Photorhabdus luminescens (バクテリア) |
| 分子量 | 理論値: 1.5 MDa |
-分子 #1: TcdA1
| 分子 | 名称: TcdA1 / タイプ: protein_or_peptide / ID: 1 / コピー数: 5 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Photorhabdus luminescens (バクテリア) Photorhabdus luminescens (バクテリア) |
| 分子量 | 理論値: 285.489844 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MAHHHHHHSS GLEVLFQGPM NESVKEIPDV LKSQCGFNCL TDISHSSFNE FRQQVSEHLS WSETHDLYHD AQQAQKDNRL YEARILKRA NPQLQNAVHL AILAPNAELI GYNNQFSGRA SQYVAPGTVS SMFSPAAYLT ELYREARNLH ASDSVYYLDT R RPDLKSMA ...文字列: MAHHHHHHSS GLEVLFQGPM NESVKEIPDV LKSQCGFNCL TDISHSSFNE FRQQVSEHLS WSETHDLYHD AQQAQKDNRL YEARILKRA NPQLQNAVHL AILAPNAELI GYNNQFSGRA SQYVAPGTVS SMFSPAAYLT ELYREARNLH ASDSVYYLDT R RPDLKSMA LSQQNMDIEL STLSLSNELL LESIKTESKL ENYTKVMEML STFRPSGATP YHDAYENVRE VIQLQDPGLE QL NASPAIA GLMHQASLLG INASISPELF NILTEEITEG NAEELYKKNF GNIEPASLAM PEYLKRYYNL SDEELSQFIG KAS NFGQQE YSNNQLITPV VNSSDGTVKV YRITREYTTN AYQMDVELFP FGGENYRLDY KFKNFYNASY LSIKLNDKRE LVRT EGAPQ VNIEYSANIT LNTADISQPF EIGLTRVLPS GSWAYAAAKF TVEEYNQYSF LLKLNKAIRL SRATELSPTI LEGIV RSVN LQLDINTDVL GKVFLTKYYM QRYAIHAETA LILCNAPISQ RSYDNQPSQF DRLFNTPLLN GQYFSTGDEE IDLNSG STG DWRKTILKRA FNIDDVSLFR LLWITDHDNK DGKIKNNLKN LSNLYIGKLL ADIHQLTIDE LDLLLIAVGE GKTNLSA IS DKQLATLIRK LNTITSWLHT QKWSVFQLFI MTSTSYNKTL TPEIKNLLDT VYHGLQGFDK DKADLLHVMA PYIAATLQ L SSENVAHSVL LWADKLQPGD GAMTAEKFWD WLNTKYTPGS SEAVETQEHI VQYCQALAQL EMVYHSTGIN ENAFRLFVT KPEMFGAATG AAPAHDALSL IMLTRFADWV NALGEKASSV LAAFEANSLT AEQLADAMNL DANLLLQASI QAQNHQHLPP VTPENAFSC WTSINTILQW VNVAQQLNVA PQGVSALVGL DYIQSMKETP TYAQWENAAG VLTAGLNSQQ ANTLHAFLDE S RSAALSTY YIRQVAKAAA AIKSRDDLYQ YLLIDNQVSA AIKTTRIAEA IASIQLYVNR ALENVEENAN SGVISRQFFI DW DKYNKRY STWAGVSQLV YYPENYIDPT MRIGQTKMMD ALLQSVSQSQ LNADTVEDAF MSYLTSFEQV ANLKVISAYH DNI NNDQGL TYFIGLSETD AGEYYWRSVD HSKFNDGKFA ANAWSEWHKI DCPINPYKST IRPVIYKSRL YLLWLEQKEI TKQT GNSKD GYQTETDYRY ELKLAHIRYD GTWNTPITFD VNKKISELKL EKNRAPGLYC AGYQGEDTLL VMFYNQQDTL DSYKN ASMQ GLYIFADMAS KDMTPEQSNV YRDNSYQQFD TNNVRRVNNR YAEDYEIPSS VSSRKDYGWG DYYLSMVYNG DIPTIN YKA ASSDLKIYIS PKLRIIHNGY EGQKRNQCNL MNKYGKLGDK FIVYTSLGVN PNNSSNKLMF YPVYQYSGNT SGLNQGR LL FHRDTTYPSK VEAWIPGAKR SLTNQNAAIG DDYATDSLNK PDDLKQYIFM TDSKGTATDV SGPVEINTAI SPAKVQII V KAGGKEQTFT ADKDVSIQPS PSFDEMNYQF NALEIDGSGL NFINNSASID VTFTAFAEDG RKLGYESFSI PVTLKVSTD NALTLHHNEN GAQYMQWQSY RTRLNTLFAR QLVARATTGI DTILSMETQN IQEPQLGKGF YATFVIPPYN LSTHGDERWF KLYIKHVVD NNSHIIYSGQ LTDTNINITL FIPLDDVPLN QDYHAKVYMT FKKSPSDGTW WGPHFVRDDK GIVTINPKSI L THFESVNV LNNISSEPMD FSGANSLYFW ELFYYTPMLV AQRLLHEQNF DEANRWLKYV WSPSGYIVHG QIQNYQWNVR PL LEDTSWN SDPLDSVDPD AVAQHDPMHY KVSTFMRTLD LLIARGDHAY RQLERDTLNE AKMWYMQALH LLGDKPYLPL STT WSDPRL DRAADITTQN AHDSAIVALR QNIPTPAPLS LRSANTLTDL FLPQINEVMM NYWQTLAQRV YNLRHNLSID GQPL YLPIY ATPADPKALL SAAVATSQGG GWLPESFMSL WRFPHMLENA RGMVSQLTQF GSTLQNIIER QDAEALNALL QNQAA ELIL TNLSIQDKTI EELDAEKTVL EKSKAGAQSR FDSYGKLYDE NINAGENQAM TLRASAAGLT TAVQASRLAG AAADLV PNI FGFAGGGSRW GAIAEATGYV MEFSANVMNT EADKISQSET YRRRRQEWEI QRNNAEAELK QIDAQLKSLA VRREAAV LQ KTSLKTQQEQ TQSQLAFLQR KFSNQALYNW LRGRLAAIYF QFYDLAVARC LMAEQAYRWE LNDDSARFIK PGAWQGTY A GLLAGETLML SLAQMEDAHL KRDKRALEVE RTVSLAEVYA GLPKDNGPFS LAQEIDKLVS QGSGSAGSGN NNLAFGAGT DTKTSLQASV SFADLKIRED YPASLGKIRR IKQISVTLPA LLGPYQDVQA ILSYGDKAGL ANGCEALAVS HGMNDSGQFQ LDFNDGKFL PFEGIAIDQG TLTLSFPNAS MPEKGKQATM LKTLNDIILH IRYTIK UniProtKB: TcdA1 |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 11.2 構成要素:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| グリッド | モデル: Quantifoil R2/1 / 材質: COPPER / メッシュ: 300 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: CONTINUOUS / 支持フィルム - Film thickness: 2 / 前処理 - タイプ: GLOW DISCHARGE | ||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 286 K / 装置: FEI VITROBOT MARK IV |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 検出モード: INTEGRATING / 撮影したグリッド数: 2 / 実像数: 24611 / 平均電子線量: 57.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: SPOT SCAN / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 3.0 µm / 最小 デフォーカス(公称値): 1.5 µm / 倍率(公称値): 81000 |
| 試料ステージ | ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 精密化 | 空間: REAL / プロトコル: FLEXIBLE FIT |
|---|---|
| 得られたモデル |  PDB-8cq0: |
 ムービー
ムービー コントローラー
コントローラー








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)