+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1414 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Molecular architecture of the prolate head of bacteriophage T4. | |||||||||
 Map data Map data | Cryo-EM reconstruction of the bacteriophage T4 prolate head using D5 symmetry | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 21.0 Å | |||||||||
 Authors Authors | Fokine A / Chipman P / Leiman P / Mesyanzhinov V / Rao V / Rossmann M | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2004 Journal: Proc Natl Acad Sci U S A / Year: 2004Title: Molecular architecture of the prolate head of bacteriophage T4. Authors: Andrei Fokine / Paul R Chipman / Petr G Leiman / Vadim V Mesyanzhinov / Venigalla B Rao / Michael G Rossmann /  Abstract: The head of bacteriophage T4 is a prolate icosahedron with one unique portal vertex to which the phage tail is attached. The three-dimensional structure of mature bacteriophage T4 head has been ...The head of bacteriophage T4 is a prolate icosahedron with one unique portal vertex to which the phage tail is attached. The three-dimensional structure of mature bacteriophage T4 head has been determined to 22-A resolution by using cryo-electron microscopy. The T4 capsid has a hexagonal surface lattice characterized by the triangulation numbers T(end) = 13 laevo for the icosahedral caps and T(mid) = 20 for the midsection. Hexamers of the major capsid protein gene product (gp)23* and pentamers of the vertex protein gp24*, as well as the outer surface proteins highly antigenic outer capsid protein (hoc) and small outer capsid protein (soc), are clearly evident in the reconstruction. The size and shape of the gp23* hexamers are similar to the major capsid protein organization of bacteriophage HK97. The binding sites and shape of the hoc and soc proteins have been established by analysis of the soc(-) and hoc(-)soc(-) T4 structures. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1414.map.gz emd_1414.map.gz | 14.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1414-v30.xml emd-1414-v30.xml emd-1414.xml emd-1414.xml | 9.1 KB 9.1 KB | Display Display |  EMDB header EMDB header |
| Images |  1414.gif 1414.gif | 190.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1414 http://ftp.pdbj.org/pub/emdb/structures/EMD-1414 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1414 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1414 | HTTPS FTP |
-Validation report
| Summary document |  emd_1414_validation.pdf.gz emd_1414_validation.pdf.gz | 291.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1414_full_validation.pdf.gz emd_1414_full_validation.pdf.gz | 290.4 KB | Display | |
| Data in XML |  emd_1414_validation.xml.gz emd_1414_validation.xml.gz | 6.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1414 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1414 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1414 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1414 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1414.map.gz / Format: CCP4 / Size: 41.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1414.map.gz / Format: CCP4 / Size: 41.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM reconstruction of the bacteriophage T4 prolate head using D5 symmetry | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.96 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
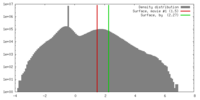
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bacteriophage T4 capsid
| Entire | Name: Bacteriophage T4 capsid |
|---|---|
| Components |
|
-Supramolecule #1000: Bacteriophage T4 capsid
| Supramolecule | Name: Bacteriophage T4 capsid / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Enterobacteria phage T4
| Supramolecule | Name: Enterobacteria phage T4 / type: virus / ID: 1 / Name.synonym: T4 capsid / NCBI-ID: 10665 / Sci species name: Enterobacteria phage T4 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No / Syn species name: T4 capsid |
|---|---|
| Host (natural) | Organism:  |
| Virus shell | Shell ID: 1 / Name: gp23 / Diameter: 1200 Å / T number (triangulation number): 13 |
| Virus shell | Shell ID: 2 / Name: gp23 / T number (triangulation number): 20 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.2 Details: Distilled water. The concentration of the virus particles was 10E11 particles per ml. |
|---|---|
| Grid | Details: quantifoil copper grid 2um holes |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 90 K / Instrument: HOMEMADE PLUNGER Details: Vitrification instrument: in-house, gravity driven plunger Method: 3.5ul of sample hand blotted approx. 1sec |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM300FEG/T |
|---|---|
| Temperature | Average: 97 K |
| Alignment procedure | Legacy - Astigmatism: live fft at 190,000x |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 100 / Average electron dose: 20 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 47000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 45000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: D5 (2x5 fold dihedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 21.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SPIDER, EMAN Details: The reconstruction was calculated assuming D5 symmetry. FSC=0.5 at 21 A, FSC=0.3 at 17 A. Number images used: 6390 |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)