[日本語] English
 万見
万見- EMDB-12003: CryoEM structure of the human sodium proton exchanger NHA2 in nanodisc -
+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-12003 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | CryoEM structure of the human sodium proton exchanger NHA2 in nanodisc | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | secondary active transporter / membrane transport / ion antiport / pH regulation / MEMBRANE PROTEIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報lithium:proton antiporter activity / lithium ion transport / positive regulation of osteoclast development / sodium ion homeostasis / sodium:proton antiporter activity / regulation of insulin secretion involved in cellular response to glucose stimulus / clathrin-dependent endocytosis / flagellated sperm motility / : / sodium ion transport ...lithium:proton antiporter activity / lithium ion transport / positive regulation of osteoclast development / sodium ion homeostasis / sodium:proton antiporter activity / regulation of insulin secretion involved in cellular response to glucose stimulus / clathrin-dependent endocytosis / flagellated sperm motility / : / sodium ion transport / recycling endosome / mitochondrial membrane / Stimuli-sensing channels / recycling endosome membrane / synaptic vesicle membrane / sperm principal piece / monoatomic ion transmembrane transport / basolateral plasma membrane / apical plasma membrane / mitochondrial inner membrane / mitochondrion / identical protein binding / plasma membrane 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
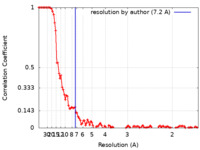
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 7.2 Å | |||||||||
 データ登録者 データ登録者 | Woehlert D / Kuhlbrandt W | |||||||||
| 資金援助 |  ドイツ, 1件 ドイツ, 1件
| |||||||||
 引用 引用 |  ジャーナル: To Be Published ジャーナル: To Be Publishedタイトル: CryoEM structure of the human sodium proton exchanger NHA2 in nanodisc 著者: Woehlert D / Kuhlbrandt W / Yildiz O | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_12003.map.gz emd_12003.map.gz | 100.3 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-12003-v30.xml emd-12003-v30.xml emd-12003.xml emd-12003.xml | 13.4 KB 13.4 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_12003_fsc.xml emd_12003_fsc.xml | 10.6 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_12003.png emd_12003.png | 44.7 KB | ||
| マスクデータ |  emd_12003_msk_1.map emd_12003_msk_1.map | 107.2 MB |  マスクマップ マスクマップ | |
| Filedesc metadata |  emd-12003.cif.gz emd-12003.cif.gz | 5.3 KB | ||
| その他 |  emd_12003_half_map_1.map.gz emd_12003_half_map_1.map.gz emd_12003_half_map_2.map.gz emd_12003_half_map_2.map.gz | 99.2 MB 99.2 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12003 http://ftp.pdbj.org/pub/emdb/structures/EMD-12003 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12003 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12003 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_12003.map.gz / 形式: CCP4 / 大きさ: 107.2 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_12003.map.gz / 形式: CCP4 / 大きさ: 107.2 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.833 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-マスク #1
| ファイル |  emd_12003_msk_1.map emd_12003_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_12003_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_12003_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Human sodium proton exchanger NHA2 in nanodisc
| 全体 | 名称: Human sodium proton exchanger NHA2 in nanodisc |
|---|---|
| 要素 |
|
-超分子 #1: Human sodium proton exchanger NHA2 in nanodisc
| 超分子 | 名称: Human sodium proton exchanger NHA2 in nanodisc / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 150 KDa |
-分子 #1: Sodium/hydrogen exchanger 9B2
| 分子 | 名称: Sodium/hydrogen exchanger 9B2 / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 57.611672 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MGDEDKRITY EDSEPSTGMN YTPSMHQEAQ EETVMKLKGI DANEPTEGSI LLKSSEKKLQ ETPTEANHVQ RLRQMLACPP HGLLDRVIT NVTIIVLLWA VVWSITGSEC LPGGNLFGII ILFYCAIIGG KLLGLIKLPT LPPLPSLLGM LLAGFLIRNI P VINDNVQI ...文字列: MGDEDKRITY EDSEPSTGMN YTPSMHQEAQ EETVMKLKGI DANEPTEGSI LLKSSEKKLQ ETPTEANHVQ RLRQMLACPP HGLLDRVIT NVTIIVLLWA VVWSITGSEC LPGGNLFGII ILFYCAIIGG KLLGLIKLPT LPPLPSLLGM LLAGFLIRNI P VINDNVQI KHKWSSSLRS IALSIILVRA GLGLDSKALK KLKGVCVRLS MGPCIVEACT SALLAHYLLG LPWQWGFILG FV LGAVSPA VVVPSMLLLQ GGGYGVEKGV PTLLMAAGSF DDILAITGFN TCLGIAFSTG STVFNVLRGV LEVVIGVATG SVL GFFIQY FPSRDQDKLV CKRTFLVLGL SVLAVFSSVH FGFPGSGGLC TLVMAFLAGM GWTSEKAEVE KIIAVAWDIF QPLL FGLIG AEVSIASLRP ETVGLCVATV GIAVLIRILT TFLMVCFAGF NLKEKIFISF AWLPKATVQA AIGSVALDTA RSHGE KQLE DYGMDVLTVA FLSILITAPI GSLLIGLLGP RLLQKVEHQN KDEEVQGETS VQV UniProtKB: Sodium/hydrogen exchanger 9B2 |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 3 mg/mL |
|---|---|
| 緩衝液 | pH: 7.4 |
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 実像数: 1251 / 平均電子線量: 40.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 精密化 | 空間: REAL / プロトコル: AB INITIO MODEL |
|---|---|
| 得られたモデル |  PDB-7b4m: |
 ムービー
ムービー コントローラー
コントローラー













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)