[English] 日本語
 Yorodumi
Yorodumi- EMDB-11872: Cryo-EM structure of Arabidopsis thaliana Complex-I (membrane core) -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11872 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Arabidopsis thaliana Complex-I (membrane core) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex-I / ELECTRON TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationanther dehiscence / vegetative to reproductive phase transition of meristem / Lyases; Carbon-oxygen lyases; Hydro-lyases / P450-containing electron transport chain / photorespiration / embryo development ending in seed dormancy / NADH dehydrogenase complex / response to abscisic acid / plant-type vacuole / cellular respiration ...anther dehiscence / vegetative to reproductive phase transition of meristem / Lyases; Carbon-oxygen lyases; Hydro-lyases / P450-containing electron transport chain / photorespiration / embryo development ending in seed dormancy / NADH dehydrogenase complex / response to abscisic acid / plant-type vacuole / cellular respiration / regulation of reactive oxygen species metabolic process / plastid / protein homotrimerization / NADH:ubiquinone reductase (H+-translocating) / mitochondrial electron transport, NADH to ubiquinone / mitochondrial respiratory chain complex I assembly / NADH dehydrogenase activity / oxidoreductase activity, acting on NAD(P)H / respiratory chain complex I / NADH dehydrogenase (ubiquinone) activity / ATP synthesis coupled electron transport / response to salt stress / carbonate dehydratase activity / respiratory electron transport chain / mitochondrial intermembrane space / 2 iron, 2 sulfur cluster binding / mitochondrial membrane / peroxisome / mitochondrial inner membrane / copper ion binding / nucleolus / mitochondrion / extracellular region / metal ion binding / identical protein binding / nucleus / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
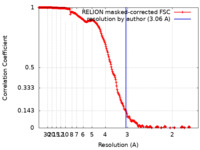
| Method | single particle reconstruction / cryo EM / Resolution: 3.06 Å | |||||||||
 Authors Authors | Klusch N / Kuehlbrandt W | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Plant Cell / Year: 2021 Journal: Plant Cell / Year: 2021Title: A ferredoxin bridge connects the two arms of plant mitochondrial complex I. Authors: Niklas Klusch / Jennifer Senkler / Özkan Yildiz / Werner Kühlbrandt / Hans-Peter Braun /  Abstract: Mitochondrial complex I is the main site for electron transfer to the respiratory chain and generates much of the proton gradient across the inner mitochondrial membrane. Complex I is composed of two ...Mitochondrial complex I is the main site for electron transfer to the respiratory chain and generates much of the proton gradient across the inner mitochondrial membrane. Complex I is composed of two arms, which form a conserved L-shape. We report the structures of the intact, 47-subunit mitochondrial complex I from Arabidopsis thaliana and the 51-subunit complex I from the green alga Polytomella sp., both at around 2.9 Å resolution. In both complexes, a heterotrimeric γ-carbonic anhydrase domain is attached to the membrane arm on the matrix side. Two states are resolved in A. thaliana complex I, with different angles between the two arms and different conformations of the ND1 (NADH dehydrogenase subunit 1) loop near the quinol binding site. The angle appears to depend on a bridge domain, which links the peripheral arm to the membrane arm and includes an unusual ferredoxin. We propose that the bridge domain participates in regulating the activity of plant complex I. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11872.map.gz emd_11872.map.gz | 40.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11872-v30.xml emd-11872-v30.xml emd-11872.xml emd-11872.xml | 39.9 KB 39.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11872_fsc.xml emd_11872_fsc.xml | 21.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_11872.png emd_11872.png | 141.2 KB | ||
| Filedesc metadata |  emd-11872.cif.gz emd-11872.cif.gz | 9.8 KB | ||
| Others |  emd_11872_half_map_1.map.gz emd_11872_half_map_1.map.gz emd_11872_half_map_2.map.gz emd_11872_half_map_2.map.gz | 631.2 MB 631.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11872 http://ftp.pdbj.org/pub/emdb/structures/EMD-11872 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11872 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11872 | HTTPS FTP |
-Related structure data
| Related structure data |  7aqqMC  7aqrC  7aqwC  7ar7C  7ar8C  7ar9C  7arbC  7arcC  7ardC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11872.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11872.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.837 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
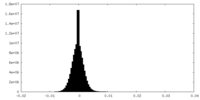
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: #2
| File | emd_11872_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
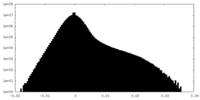
| Density Histograms |
-Half map: #1
| File | emd_11872_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
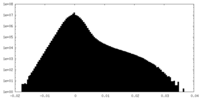
| Density Histograms |
- Sample components
Sample components
+Entire : Arabidopsis complex I - membrane core
+Supramolecule #1: Arabidopsis complex I - membrane core
+Macromolecule #1: NADH-ubiquinone oxidoreductase chain 3
+Macromolecule #2: NADH-ubiquinone oxidoreductase chain 1
+Macromolecule #3: NADH-ubiquinone oxidoreductase chain 6
+Macromolecule #4: NADH dehydrogenase subunit 4L
+Macromolecule #5: NADH-ubiquinone oxidoreductase chain 5
+Macromolecule #6: NADH-ubiquinone oxidoreductase chain 4
+Macromolecule #7: NADH-ubiquinone oxidoreductase chain 2
+Macromolecule #8: AT3G07480.1
+Macromolecule #9: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 8-B
+Macromolecule #10: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 13-A
+Macromolecule #11: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 1
+Macromolecule #12: At2g46540/F11C10.23
+Macromolecule #13: Excitatory amino acid transporter
+Macromolecule #14: NADH dehydrogenase [ubiquinone] iron-sulfur protein 5-B
+Macromolecule #15: At4g16450
+Macromolecule #16: P1
+Macromolecule #17: unknown
+Macromolecule #18: Uncharacterized protein At2g27730, mitochondrial
+Macromolecule #19: Gamma carbonic anhydrase-like 2, mitochondrial
+Macromolecule #20: Gamma carbonic anhydrase 2, mitochondrial
+Macromolecule #21: Gamma carbonic anhydrase 1, mitochondrial
+Macromolecule #22: Ubiquinone-9
+Macromolecule #23: PHOSPHATIDYLETHANOLAMINE
+Macromolecule #24: Lauryl Maltose Neopentyl Glycol
+Macromolecule #25: FE (III) ION
+Macromolecule #26: (7S)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5...
+Macromolecule #27: ZINC ION
+Macromolecule #28: (1S)-2-{[{[(2R)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-...
+Macromolecule #29: 1,2-DICAPROYL-SN-PHOSPHATIDYL-L-SERINE
+Macromolecule #30: Phosphatidylinositol
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 43.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)