[English] 日本語
 Yorodumi
Yorodumi- EMDB-11287: Human mitochondrial ribosome in complex with mRNA, A/A tRNA and P... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11287 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human mitochondrial ribosome in complex with mRNA, A/A tRNA and P/P tRNA, local-masked aligned on SSU head. | |||||||||
 Map data Map data | Sharpened, local-resolution filtered and masked. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
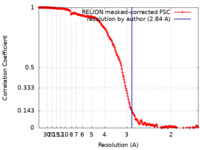
| Method | single particle reconstruction / cryo EM / Resolution: 2.84 Å | |||||||||
 Authors Authors | Andrell A / Itoh Y / Amunts A | |||||||||
| Funding support |  Sweden, 1 items Sweden, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2021 Journal: Science / Year: 2021Title: Mechanism of membrane-tethered mitochondrial protein synthesis. Authors: Yuzuru Itoh / Juni Andréll / Austin Choi / Uwe Richter / Priyanka Maiti / Robert B Best / Antoni Barrientos / Brendan J Battersby / Alexey Amunts /     Abstract: Mitochondrial ribosomes (mitoribosomes) are tethered to the mitochondrial inner membrane to facilitate the cotranslational membrane insertion of the synthesized proteins. We report cryo-electron ...Mitochondrial ribosomes (mitoribosomes) are tethered to the mitochondrial inner membrane to facilitate the cotranslational membrane insertion of the synthesized proteins. We report cryo-electron microscopy structures of human mitoribosomes with nascent polypeptide, bound to the insertase oxidase assembly 1-like (OXA1L) through three distinct contact sites. OXA1L binding is correlated with a series of conformational changes in the mitoribosomal large subunit that catalyze the delivery of newly synthesized polypeptides. The mechanism relies on the folding of mL45 inside the exit tunnel, forming two specific constriction sites that would limit helix formation of the nascent chain. A gap is formed between the exit and the membrane, making the newly synthesized proteins accessible. Our data elucidate the basis by which mitoribosomes interact with the OXA1L insertase to couple protein synthesis and membrane delivery. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11287.map.gz emd_11287.map.gz | 14.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11287-v30.xml emd-11287-v30.xml emd-11287.xml emd-11287.xml | 37.7 KB 37.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11287_fsc.xml emd_11287_fsc.xml | 19.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_11287.png emd_11287.png | 72.4 KB | ||
| Masks |  emd_11287_msk_1.map emd_11287_msk_1.map | 600.7 MB |  Mask map Mask map | |
| Others |  emd_11287_half_map_1.map.gz emd_11287_half_map_1.map.gz emd_11287_half_map_2.map.gz emd_11287_half_map_2.map.gz | 486.3 MB 486.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11287 http://ftp.pdbj.org/pub/emdb/structures/EMD-11287 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11287 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11287 | HTTPS FTP |
-Validation report
| Summary document |  emd_11287_validation.pdf.gz emd_11287_validation.pdf.gz | 399.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11287_full_validation.pdf.gz emd_11287_full_validation.pdf.gz | 398.5 KB | Display | |
| Data in XML |  emd_11287_validation.xml.gz emd_11287_validation.xml.gz | 496 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11287 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11287 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11287 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11287 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11287.map.gz / Format: CCP4 / Size: 39.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11287.map.gz / Format: CCP4 / Size: 39.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened, local-resolution filtered and masked. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
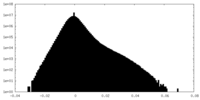
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11287_msk_1.map emd_11287_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
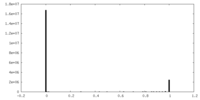
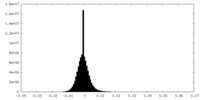
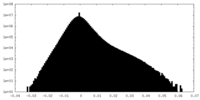
| Density Histograms |
-Half map: #2
| File | emd_11287_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
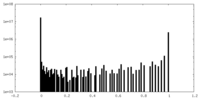
| Density Histograms |
-Half map: #1
| File | emd_11287_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
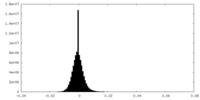
| Density Histograms |
- Sample components
Sample components
-Entire : Ribosome with mRNA and tRNAs
| Entire | Name: Ribosome with mRNA and tRNAs |
|---|---|
| Components |
|
-Supramolecule #1: Ribosome with mRNA and tRNAs
| Supramolecule | Name: Ribosome with mRNA and tRNAs / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#89 Details: Mitochondrial ribosome in complex with mRNA, A/A tRNA and P/P tRNA |
|---|---|
| Molecular weight | Theoretical: 3 MDa |
-Supramolecule #2: SSU body
| Supramolecule | Name: SSU body / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#53 / Details: The body part of the small subunit of ribosome |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 3.0 nm / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Digitization - Frames/image: 1-20 / Number grids imaged: 2 / Number real images: 19655 / Average exposure time: 4.0 sec. / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 3.6 µm / Calibrated defocus min: 0.2 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)