+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0933 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of immature Zika virus | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | immature Zika virus / virus / flavivirus | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationflavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity / negative regulation of innate immune response / viral capsid / double-stranded RNA binding / nucleoside-triphosphate phosphatase / 4 iron, 4 sulfur cluster binding / clathrin-dependent endocytosis of virus by host cell ...flavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity / negative regulation of innate immune response / viral capsid / double-stranded RNA binding / nucleoside-triphosphate phosphatase / 4 iron, 4 sulfur cluster binding / clathrin-dependent endocytosis of virus by host cell / molecular adaptor activity / mRNA (guanine-N7)-methyltransferase / methyltransferase cap1 / methyltransferase cap1 activity / mRNA 5'-cap (guanine-N7-)-methyltransferase activity / RNA helicase activity / protein dimerization activity / host cell perinuclear region of cytoplasm / host cell endoplasmic reticulum membrane / RNA helicase / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / serine-type endopeptidase activity / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / viral RNA genome replication / RNA-directed RNA polymerase activity / fusion of virus membrane with host endosome membrane / viral envelope / centrosome / lipid binding / virion attachment to host cell / GTP binding / host cell nucleus / virion membrane / structural molecule activity / ATP hydrolysis activity / proteolysis / extracellular region / ATP binding / metal ion binding Similarity search - Function | ||||||||||||
| Biological species |  Zika virus (isolate ZIKV/Human/French Polynesia/10087PF/2013) / Zika virus (isolate ZIKV/Human/French Polynesia/10087PF/2013) /  Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 9.0 Å | ||||||||||||
 Authors Authors | Tan TY / Fibriansah G | ||||||||||||
| Funding support |  Singapore, 3 items Singapore, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Capsid protein structure in Zika virus reveals the flavivirus assembly process. Authors: Ter Yong Tan / Guntur Fibriansah / Victor A Kostyuchenko / Thiam-Seng Ng / Xin-Xiang Lim / Shuijun Zhang / Xin-Ni Lim / Jiaqi Wang / Jian Shi / Marc C Morais / Davide Corti / Shee-Mei Lok /    Abstract: Structures of flavivirus (dengue virus and Zika virus) particles are known to near-atomic resolution and show detailed structure and arrangement of their surface proteins (E and prM in immature virus ...Structures of flavivirus (dengue virus and Zika virus) particles are known to near-atomic resolution and show detailed structure and arrangement of their surface proteins (E and prM in immature virus or M in mature virus). By contrast, the arrangement of the capsid proteins:RNA complex, which forms the core of the particle, is poorly understood, likely due to inherent dynamics. Here, we stabilize immature Zika virus via an antibody that binds across the E and prM proteins, resulting in a subnanometer resolution structure of capsid proteins within the virus particle. Fitting of the capsid protein into densities shows the presence of a helix previously thought to be removed via proteolysis. This structure illuminates capsid protein quaternary organization, including its orientation relative to the lipid membrane and the genomic RNA, and its interactions with the transmembrane regions of the surface proteins. Results show the capsid protein plays a central role in the flavivirus assembly process. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0933.map.gz emd_0933.map.gz | 522.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0933-v30.xml emd-0933-v30.xml emd-0933.xml emd-0933.xml | 13.3 KB 13.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0933.png emd_0933.png | 230.7 KB | ||
| Filedesc metadata |  emd-0933.cif.gz emd-0933.cif.gz | 5.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0933 http://ftp.pdbj.org/pub/emdb/structures/EMD-0933 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0933 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0933 | HTTPS FTP |
-Related structure data
| Related structure data |  6lnuMC  0932C  0934C  6lntC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0933.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0933.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
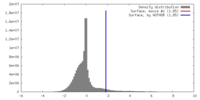
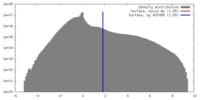
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013
| Entire | Name:  Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 |
|---|---|
| Components |
|
-Supramolecule #1: Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013
| Supramolecule | Name: Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 type: virus / ID: 1 / Parent: 0 / Macromolecule list: all Details: The virus was isolated from Zika patient. The immature Zika virus was grown in Aedes Albopictus clone C6/36 cell. NCBI-ID: 2043570 Sci species name: Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 Sci species strain: H/PF/2013 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|
-Macromolecule #1: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO / EC number: flavivirin |
|---|---|
| Source (natural) | Organism:  Zika virus (isolate ZIKV/Human/French Polynesia/10087PF/2013) Zika virus (isolate ZIKV/Human/French Polynesia/10087PF/2013)Strain: isolate ZIKV/Human/French Polynesia/10087PF/2013 |
| Molecular weight | Theoretical: 54.444051 KDa |
| Sequence | String: IRCIGVSNRD FVEGMSGGTW VDVVLEHGGC VTVMAQDKPT VDIELVTTTV SNMAEVRSYC YEASISDMAS DSRCPTQGEA YLDKQSDTQ YVCKRTLVDR GWGNGCGLFG KGSLVTCAKF ACSKKMTGKS IQPENLEYRI MLSVHGSQHS GMIVNDTGHE T DENRAKVE ...String: IRCIGVSNRD FVEGMSGGTW VDVVLEHGGC VTVMAQDKPT VDIELVTTTV SNMAEVRSYC YEASISDMAS DSRCPTQGEA YLDKQSDTQ YVCKRTLVDR GWGNGCGLFG KGSLVTCAKF ACSKKMTGKS IQPENLEYRI MLSVHGSQHS GMIVNDTGHE T DENRAKVE ITPNSPRAEA TLGGFGSLGL DCEPRTGLDF SDLYYLTMNN KHWLVHKEWF HDIPLPWHAG ADTGTPHWNN KE ALVEFKD AHAKRQTVVV LGSQEGAVHT ALAGALEAEM DGAKGRLSSG HLKCRLKMDK LRLKGVSYSL CTAAFTFTKI PAE TLHGTV TVEVQYAGTD GPCKVPAQMA VDMQTLTPVG RLITANPVIT ESTENSKMML ELDPPFGDSY IVIGVGEKKI THHW HRSGS TIGKAFEATV RGAKRMAVLG DTAWDFGSVG GALNSLGKGI HQIFGAAFKS LFGGMSWFSQ ILIGTLLMWL GLNTK NGSI SLMCLALGGV LIFLSTAVSA UniProtKB: Genome polyprotein |
-Macromolecule #2: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO / EC number: flavivirin |
|---|---|
| Source (natural) | Organism:  Zika virus (isolate ZIKV/Human/French Polynesia/10087PF/2013) Zika virus (isolate ZIKV/Human/French Polynesia/10087PF/2013)Strain: isolate ZIKV/Human/French Polynesia/10087PF/2013 |
| Molecular weight | Theoretical: 18.561266 KDa |
| Sequence | String: RGSAYYMYLD RNDAGEAISF PTTLGMNKCY IQIMDLGHMC DATMSYECPM LDEGVEPDDV DCWCNTTSTW VVYGTCHHKK GEARRSRRA VTLPSHSTRK LQTRSQTWLE SREYTKHLIR VENWIFRNPG FALAAAAIAW LLGSSTSQKV IYLVMILLIA P AYS UniProtKB: Genome polyprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Details: 10 mM Tris-HCl, 120 mM NaCl, 1 mM EDTA, pH 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
| Details | the virus sample was incubated at 37deg C for 30 min prior to sample blotting onto the grid. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 18.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 59000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Resolution.type: BY AUTHOR / Resolution: 9.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 7922 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 2.1) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 2.1) |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)