+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6w74 | ||||||
|---|---|---|---|---|---|---|---|
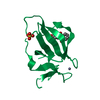
| Title | Structure of cIAP with compound 15 | ||||||
 Components Components | Baculoviral IAP repeat-containing protein 2 | ||||||
 Keywords Keywords |  LIGASE / cIAP E3 PROTAC LIGASE / cIAP E3 PROTAC | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of ripoptosome assembly involved in necroptotic process / regulation of cysteine-type endopeptidase activity / FBXO family protein binding / regulation of RIG-I signaling pathway / positive regulation of protein K48-linked ubiquitination / regulation of non-canonical NF-kappaB signal transduction / TNF receptor superfamily (TNFSF) members mediating non-canonical NF-kB pathway / regulation of necroptotic process / regulation of nucleotide-binding domain, leucine rich repeat containing receptor signaling pathway / positive regulation of protein K63-linked ubiquitination ...negative regulation of ripoptosome assembly involved in necroptotic process / regulation of cysteine-type endopeptidase activity / FBXO family protein binding / regulation of RIG-I signaling pathway / positive regulation of protein K48-linked ubiquitination / regulation of non-canonical NF-kappaB signal transduction / TNF receptor superfamily (TNFSF) members mediating non-canonical NF-kB pathway / regulation of necroptotic process / regulation of nucleotide-binding domain, leucine rich repeat containing receptor signaling pathway / positive regulation of protein K63-linked ubiquitination / CD40 receptor complex / XY body / negative regulation of necroptotic process / regulation of reactive oxygen species metabolic process / positive regulation of protein monoubiquitination / TNFR1-induced proapoptotic signaling / RIPK1-mediated regulated necrosis /  regulation of toll-like receptor signaling pathway / regulation of toll-like receptor signaling pathway /  regulation of innate immune response / Apoptotic cleavage of cellular proteins / regulation of innate immune response / Apoptotic cleavage of cellular proteins /  regulation of cell differentiation / non-canonical NF-kappaB signal transduction / necroptotic process / cysteine-type endopeptidase inhibitor activity involved in apoptotic process / canonical NF-kappaB signal transduction / response to cAMP / tumor necrosis factor-mediated signaling pathway / TICAM1, RIP1-mediated IKK complex recruitment / IKK complex recruitment mediated by RIP1 / TNFR1-induced NF-kappa-B signaling pathway / regulation of cell differentiation / non-canonical NF-kappaB signal transduction / necroptotic process / cysteine-type endopeptidase inhibitor activity involved in apoptotic process / canonical NF-kappaB signal transduction / response to cAMP / tumor necrosis factor-mediated signaling pathway / TICAM1, RIP1-mediated IKK complex recruitment / IKK complex recruitment mediated by RIP1 / TNFR1-induced NF-kappa-B signaling pathway /  ubiquitin binding / positive regulation of protein ubiquitination / TNFR2 non-canonical NF-kB pathway / placenta development / Regulation of TNFR1 signaling / RING-type E3 ubiquitin transferase / NOD1/2 Signaling Pathway / Regulation of necroptotic cell death / cytoplasmic side of plasma membrane / protein polyubiquitination / ubiquitin-protein transferase activity / ubiquitin binding / positive regulation of protein ubiquitination / TNFR2 non-canonical NF-kB pathway / placenta development / Regulation of TNFR1 signaling / RING-type E3 ubiquitin transferase / NOD1/2 Signaling Pathway / Regulation of necroptotic cell death / cytoplasmic side of plasma membrane / protein polyubiquitination / ubiquitin-protein transferase activity /  ubiquitin protein ligase activity / regulation of cell population proliferation / protein-folding chaperone binding / ubiquitin protein ligase activity / regulation of cell population proliferation / protein-folding chaperone binding /  regulation of inflammatory response / regulation of inflammatory response /  transferase activity / regulation of apoptotic process / proteasome-mediated ubiquitin-dependent protein catabolic process / positive regulation of canonical NF-kappaB signal transduction / response to ethanol / transferase activity / regulation of apoptotic process / proteasome-mediated ubiquitin-dependent protein catabolic process / positive regulation of canonical NF-kappaB signal transduction / response to ethanol /  transcription coactivator activity / cell surface receptor signaling pathway / response to hypoxia / Ub-specific processing proteases / transcription coactivator activity / cell surface receptor signaling pathway / response to hypoxia / Ub-specific processing proteases /  regulation of cell cycle / apoptotic process / protein-containing complex binding / negative regulation of apoptotic process / zinc ion binding / identical protein binding / regulation of cell cycle / apoptotic process / protein-containing complex binding / negative regulation of apoptotic process / zinc ion binding / identical protein binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.11 Å MOLECULAR REPLACEMENT / Resolution: 2.11 Å | ||||||
 Authors Authors | Calabrese, M.F. / Schiemer, J.S. | ||||||
 Citation Citation |  Journal: Nat.Chem.Biol. / Year: 2020 Journal: Nat.Chem.Biol. / Year: 2020Title: Structural Characterization of BTK:PROTAC:cIAP Ternary Complexes: From Snapshots to Ensembles Authors: Calabrese, M.F. / Schiemer, J.S. / Horst, R. / Meng, Y. / Montgomery, J. / Xu, Y. / Feng, X. / Borzilleri, K. / Uccello, D.P. / Leverett, C. / Brown, S. / Che, Y. / Brown, M.F. / Hayward, M. ...Authors: Calabrese, M.F. / Schiemer, J.S. / Horst, R. / Meng, Y. / Montgomery, J. / Xu, Y. / Feng, X. / Borzilleri, K. / Uccello, D.P. / Leverett, C. / Brown, S. / Che, Y. / Brown, M.F. / Hayward, M.M. / Gilbert, A.M. / Noe, M.C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6w74.cif.gz 6w74.cif.gz | 57.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6w74.ent.gz pdb6w74.ent.gz | 36.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6w74.json.gz 6w74.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/w7/6w74 https://data.pdbj.org/pub/pdb/validation_reports/w7/6w74 ftp://data.pdbj.org/pub/pdb/validation_reports/w7/6w74 ftp://data.pdbj.org/pub/pdb/validation_reports/w7/6w74 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6w7oC  6w8iC  4kmnS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 11318.731 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: BIRC2, API1, MIHB, RNF48 / Production host: Homo sapiens (human) / Gene: BIRC2, API1, MIHB, RNF48 / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli)References: UniProt: Q13490, RING-type E3 ubiquitin transferase |
|---|---|
| #2: Chemical | ChemComp-ZN / |
| #3: Chemical | ChemComp-TKY / |
| #4: Water | ChemComp-HOH /  Water Water |
| Has ligand of interest | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.02 Å3/Da / Density % sol: 39.15 % |
|---|---|
Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop Details: 0.2M Magnesium Acetate Tetrahydrate, 20% w/v PEG3350 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X06SA / Wavelength: 0.99988 Å / Beamline: X06SA / Wavelength: 0.99988 Å |
| Detector | Type: DECTRIS EIGER X 16M / Detector: PIXEL / Date: Feb 3, 2019 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.99988 Å / Relative weight: 1 : 0.99988 Å / Relative weight: 1 |
| Reflection | Resolution: 2.1→27.4 Å / Num. obs: 4750 / % possible obs: 88.6 % / Redundancy: 3.3 % / Biso Wilson estimate: 34.15 Å2 / CC1/2: 0.997 / Rmerge(I) obs: 0.07 / Net I/σ(I): 9.9 |
| Reflection shell | Resolution: 2.107→2.116 Å / Rmerge(I) obs: 0.418 / Mean I/σ(I) obs: 2.3 / Num. unique obs: 63 / CC1/2: 0.96 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 4kmn Resolution: 2.11→27.4 Å / Cor.coef. Fo:Fc: 0.943 / Cor.coef. Fo:Fc free: 0.896 / SU R Cruickshank DPI: 0.281 / Cross valid method: THROUGHOUT / σ(F): 0 / SU R Blow DPI: 0.306 / SU Rfree Blow DPI: 0.232 / SU Rfree Cruickshank DPI: 0.228
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 90.23 Å2 / Biso mean: 43.34 Å2 / Biso min: 23.05 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.29 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.11→27.4 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.11→2.16 Å / Rfactor Rfree error: 0 / Total num. of bins used: 12
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 58.2869 Å / Origin y: 5.6017 Å / Origin z: 12.7624 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: { A|* } |
 Movie
Movie Controller
Controller












 PDBj
PDBj














