[English] 日本語
 Yorodumi
Yorodumi- PDB-4uz3: Crystal structure of the N-terminal LysM domains from the putativ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4uz3 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal structure of the N-terminal LysM domains from the putative NlpC/P60 D,L endopeptidase from T. thermophilus bound to N-acetyl-chitohexaose | |||||||||
 Components Components | CELL WALL-BINDING ENDOPEPTIDASE-RELATED PROTEIN | |||||||||
 Keywords Keywords |  HYDROLASE HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationMembrane-bound Lytic Murein Transglycosylase D; Chain A /  LysM domain / NlpC/P60 domain profile. / LysM domain / NlpC/P60 domain profile. /  Endopeptidase, NLPC/P60 domain / NlpC/P60 family / Lysin motif / LysM domain superfamily / Endopeptidase, NLPC/P60 domain / NlpC/P60 family / Lysin motif / LysM domain superfamily /  LysM domain / LysM domain profile. / LysM domain / LysM domain profile. /  LysM domain ...Membrane-bound Lytic Murein Transglycosylase D; Chain A / LysM domain ...Membrane-bound Lytic Murein Transglycosylase D; Chain A /  LysM domain / NlpC/P60 domain profile. / LysM domain / NlpC/P60 domain profile. /  Endopeptidase, NLPC/P60 domain / NlpC/P60 family / Lysin motif / LysM domain superfamily / Endopeptidase, NLPC/P60 domain / NlpC/P60 family / Lysin motif / LysM domain superfamily /  LysM domain / LysM domain profile. / LysM domain / LysM domain profile. /  LysM domain / Papain-like cysteine peptidase superfamily / Roll / Alpha Beta LysM domain / Papain-like cysteine peptidase superfamily / Roll / Alpha BetaSimilarity search - Domain/homology | |||||||||
| Biological species |    THERMUS THERMOPHILUS (bacteria) THERMUS THERMOPHILUS (bacteria) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.75 Å MOLECULAR REPLACEMENT / Resolution: 1.75 Å | |||||||||
 Authors Authors | Wong, J.E.M.M. / Blaise, M. | |||||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 2015 Journal: Acta Crystallogr.,Sect.D / Year: 2015Title: An Intermolecular Binding Mechanism Involving Multiple Lysm Domains Mediates Carbohydrate Recognition by an Endopeptidase. Authors: Wong, J.E.M.M. / Midtgaard, S.R. / Gysel, K. / Thygesen, M.B. / Sorensen, K.K. / Jensen, K.J. / Stougaard, J. / Thirup, S. / Blaise, M. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4uz3.cif.gz 4uz3.cif.gz | 127.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4uz3.ent.gz pdb4uz3.ent.gz | 102 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4uz3.json.gz 4uz3.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/uz/4uz3 https://data.pdbj.org/pub/pdb/validation_reports/uz/4uz3 ftp://data.pdbj.org/pub/pdb/validation_reports/uz/4uz3 ftp://data.pdbj.org/pub/pdb/validation_reports/uz/4uz3 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 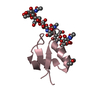
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
-Protein , 1 types, 3 molecules ABC
| #1: Protein | Mass: 10971.500 Da / Num. of mol.: 3 / Fragment: LYSM DOMAIN, RESIDUES 16-114 Source method: isolated from a genetically manipulated source Source: (gene. exp.)    THERMUS THERMOPHILUS (bacteria) / Strain: HB8 / Production host: THERMUS THERMOPHILUS (bacteria) / Strain: HB8 / Production host:   ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3) / Variant (production host): ROSETTA2 / References: UniProt: Q5SLM7 ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3) / Variant (production host): ROSETTA2 / References: UniProt: Q5SLM7 |
|---|
-Sugars , 4 types, 4 molecules 
| #2: Polysaccharide | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2- ...2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose / Mass: 1237.172 Da / Num. of mol.: 1 / Mass: 1237.172 Da / Num. of mol.: 1Source method: isolated from a genetically manipulated source |
|---|---|
| #3: Polysaccharide | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2- ...2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose / Mass: 1237.172 Da / Num. of mol.: 1 / Mass: 1237.172 Da / Num. of mol.: 1Source method: isolated from a genetically manipulated source |
| #4: Polysaccharide | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2- ...2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose / Mass: 1237.172 Da / Num. of mol.: 1 / Mass: 1237.172 Da / Num. of mol.: 1Source method: isolated from a genetically manipulated source |
| #5: Sugar | ChemComp-NDG /  N-Acetylglucosamine N-Acetylglucosamine |
-Non-polymers , 3 types, 398 molecules 




| #6: Chemical |  Sulfate Sulfate#7: Chemical |  1,4-Dioxane 1,4-Dioxane#8: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.97 Å3/Da / Density % sol: 58.61 % / Description: NONE |
|---|---|
Crystal grow | Details: 0.1 M MES PH6.5, 1.6 M AMMONIUM SULFATE, 5% DIOXANE |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  MAX II MAX II  / Beamline: I911-2 / Wavelength: 1 / Beamline: I911-2 / Wavelength: 1 |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Nov 27, 2013 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1 Å / Relative weight: 1 : 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.75→20 Å / Num. obs: 39631 / % possible obs: 99.9 % / Observed criterion σ(I): 2 / Redundancy: 12.3 % / Biso Wilson estimate: 14.78 Å2 / Rmerge(I) obs: 0.01 / Net I/σ(I): 20.3 |
| Reflection shell | Resolution: 1.75→1.8 Å / Redundancy: 12.3 % / Rmerge(I) obs: 0.87 / Mean I/σ(I) obs: 3.3 / % possible all: 100 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT / Resolution: 1.75→19.589 Å / SU ML: 0.16 / σ(F): 1.37 / Phase error: 17.3 / Stereochemistry target values: ML MOLECULAR REPLACEMENT / Resolution: 1.75→19.589 Å / SU ML: 0.16 / σ(F): 1.37 / Phase error: 17.3 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 18.7 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.75→19.589 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller














 PDBj
PDBj






