+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3kwb | ||||||
|---|---|---|---|---|---|---|---|
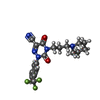
| Title | Structure of CatK covalently bound to a dioxo-triazine inhibitor | ||||||
 Components Components | Cathepsin K | ||||||
 Keywords Keywords |  HYDROLASE / HYDROLASE /  covalent bond / Cys 25 / thioimidate / Disease mutation / covalent bond / Cys 25 / thioimidate / Disease mutation /  Disulfide bond / Disulfide bond /  Glycoprotein / Glycoprotein /  Lysosome / Lysosome /  Protease / Protease /  Thiol protease / Thiol protease /  Zymogen Zymogen | ||||||
| Function / homology |  Function and homology information Function and homology information cathepsin K / mononuclear cell differentiation / cathepsin K / mononuclear cell differentiation /  intramembranous ossification / negative regulation of cartilage development / cellular response to zinc ion starvation / RUNX1 regulates transcription of genes involved in differentiation of keratinocytes / thyroid hormone generation / endolysosome lumen / Trafficking and processing of endosomal TLR / intramembranous ossification / negative regulation of cartilage development / cellular response to zinc ion starvation / RUNX1 regulates transcription of genes involved in differentiation of keratinocytes / thyroid hormone generation / endolysosome lumen / Trafficking and processing of endosomal TLR /  proteoglycan binding ... proteoglycan binding ... cathepsin K / mononuclear cell differentiation / cathepsin K / mononuclear cell differentiation /  intramembranous ossification / negative regulation of cartilage development / cellular response to zinc ion starvation / RUNX1 regulates transcription of genes involved in differentiation of keratinocytes / thyroid hormone generation / endolysosome lumen / Trafficking and processing of endosomal TLR / intramembranous ossification / negative regulation of cartilage development / cellular response to zinc ion starvation / RUNX1 regulates transcription of genes involved in differentiation of keratinocytes / thyroid hormone generation / endolysosome lumen / Trafficking and processing of endosomal TLR /  proteoglycan binding / Activation of Matrix Metalloproteinases / cysteine-type endopeptidase activator activity involved in apoptotic process / proteoglycan binding / Activation of Matrix Metalloproteinases / cysteine-type endopeptidase activator activity involved in apoptotic process /  mitophagy / Collagen degradation / mitophagy / Collagen degradation /  fibronectin binding / collagen catabolic process / extracellular matrix disassembly / cysteine-type peptidase activity / positive regulation of apoptotic signaling pathway / fibronectin binding / collagen catabolic process / extracellular matrix disassembly / cysteine-type peptidase activity / positive regulation of apoptotic signaling pathway /  bone resorption / cellular response to transforming growth factor beta stimulus / bone resorption / cellular response to transforming growth factor beta stimulus /  collagen binding / MHC class II antigen presentation / Degradation of the extracellular matrix / proteolysis involved in protein catabolic process / lysosomal lumen / response to insulin / response to organic cyclic compound / cellular response to tumor necrosis factor / response to ethanol / collagen binding / MHC class II antigen presentation / Degradation of the extracellular matrix / proteolysis involved in protein catabolic process / lysosomal lumen / response to insulin / response to organic cyclic compound / cellular response to tumor necrosis factor / response to ethanol /  lysosome / lysosome /  immune response / apical plasma membrane / external side of plasma membrane / cysteine-type endopeptidase activity / serine-type endopeptidase activity / intracellular membrane-bounded organelle / immune response / apical plasma membrane / external side of plasma membrane / cysteine-type endopeptidase activity / serine-type endopeptidase activity / intracellular membrane-bounded organelle /  proteolysis / proteolysis /  extracellular space / extracellular region / extracellular space / extracellular region /  nucleoplasm nucleoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.02 Å MOLECULAR REPLACEMENT / Resolution: 2.02 Å | ||||||
 Authors Authors | Uitdehaag, J.C.M. / van Zeeland, M. | ||||||
 Citation Citation |  Journal: Bioorg.Med.Chem.Lett. / Year: 2010 Journal: Bioorg.Med.Chem.Lett. / Year: 2010Title: Dioxo-triazines as a novel series of cathepsin K inhibitors Authors: Rankovic, Z. / Cai, J. / Fradera, X. / Dempster, M. / Mistry, A. / Mitchell, A. / Long, C. / Hamilton, E. / King, A. / Boucharens, S. / Jamieson, C. / Gillespie, J. / Cumming, I. / ...Authors: Rankovic, Z. / Cai, J. / Fradera, X. / Dempster, M. / Mistry, A. / Mitchell, A. / Long, C. / Hamilton, E. / King, A. / Boucharens, S. / Jamieson, C. / Gillespie, J. / Cumming, I. / Uitdehaag, J. / van Zeeland, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3kwb.cif.gz 3kwb.cif.gz | 105.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3kwb.ent.gz pdb3kwb.ent.gz | 84.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3kwb.json.gz 3kwb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kw/3kwb https://data.pdbj.org/pub/pdb/validation_reports/kw/3kwb ftp://data.pdbj.org/pub/pdb/validation_reports/kw/3kwb ftp://data.pdbj.org/pub/pdb/validation_reports/kw/3kwb | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  / Cathepsin O / Cathepsin X / Cathepsin O2 / Cathepsin O / Cathepsin X / Cathepsin O2Mass: 23523.480 Da / Num. of mol.: 2 / Fragment: full length / Mutation: wild type Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: CTSK, CTSO, CTSO2 / Organ (production host): ovary / Production host: Homo sapiens (human) / Gene: CTSK, CTSO, CTSO2 / Organ (production host): ovary / Production host:   Cricetulus griseus (Chinese hamster) / References: UniProt: P43235, Cricetulus griseus (Chinese hamster) / References: UniProt: P43235,  cathepsin K cathepsin K#2: Chemical | #3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.08 Å3/Da / Density % sol: 40.88 % |
|---|---|
Crystal grow | Temperature: 298 K / pH: 3.4 Details: 30%(W/V) PEG 4000 and 150 mM Ammonium Sulfate pH 3.4 3 microliter of protein at 10 mg/ml and 2 microliter of reservoir solution. , VAPOR DIFFUSION, HANGING DROP, temperature 298K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Feb 13, 2004 / Details: MIRRORS |
| Radiation | Monochromator: UNKNOWN / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.5418 Å / Relative weight: 1 : 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.017→27.461 Å / Num. obs: 23819 / % possible obs: 93.1 % / Observed criterion σ(I): 0 / Redundancy: 2.3 % / Rmerge(I) obs: 0.068 / Rsym value: 0.068 / Net I/σ(I): 13 |
| Reflection shell | Resolution: 2.01→2.12 Å / Redundancy: 2.1 % / Rmerge(I) obs: 0.181 / Mean I/σ(I) obs: 4.1 / Rsym value: 0.181 / % possible all: 82.7 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT / Resolution: 2.02→27.47 Å / Cor.coef. Fo:Fc: 0.936 / Cor.coef. Fo:Fc free: 0.908 / Occupancy max: 1 / Occupancy min: 1 / SU B: 4.329 / SU ML: 0.122 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.296 / ESU R Free: 0.194 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE MOLECULAR REPLACEMENT / Resolution: 2.02→27.47 Å / Cor.coef. Fo:Fc: 0.936 / Cor.coef. Fo:Fc free: 0.908 / Occupancy max: 1 / Occupancy min: 1 / SU B: 4.329 / SU ML: 0.122 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.296 / ESU R Free: 0.194 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 18.42 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.02→27.47 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.02→2.07 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller












 PDBj
PDBj