Entry Database : PDB / ID : 2xnzTitle xpt-pbuX C74U Riboswitch from B. subtilis bound to acetoguanamine identified by virtual screening Guanine riboswitch Keywords / / / Function / homology / / / / / Biological species BACILLUS SUBTILIS (bacteria)Method / / / Resolution : 1.59 Å Authors Daldrop, P. / Reyes, F.E. / Robinson, D.A. / Hammond, C.M. / Lilley, D.M.J. / Batey, R.T. / Brenk, R. Journal : Chem. Biol. / Year : 2011Title : Novel ligands for a purine riboswitch discovered by RNA-ligand docking.Authors : Daldrop, P. / Reyes, F.E. / Robinson, D.A. / Hammond, C.M. / Lilley, D.M. / Batey, R.T. / Brenk, R. History Deposition Aug 6, 2010 Deposition site / Processing site Revision 1.0 Apr 6, 2011 Provider / Type Revision 1.1 May 8, 2011 Group Revision 1.2 Jul 13, 2011 Group Revision 1.3 Jun 6, 2018 Group Advisory / Data collection ... Advisory / Data collection / Database references / Structure summary Category citation / citation_author ... citation / citation_author / entity / pdbx_unobs_or_zero_occ_atoms / struct Item _citation.journal_abbrev / _citation.journal_id_ISSN ... _citation.journal_abbrev / _citation.journal_id_ISSN / _citation.page_last / _citation.pdbx_database_id_DOI / _citation.title / _citation_author.name / _entity.pdbx_description / _struct.title Revision 1.4 Dec 20, 2023 Group Advisory / Data collection ... Advisory / Data collection / Database references / Derived calculations / Other / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_initial_refinement_model / pdbx_struct_conn_angle / pdbx_unobs_or_zero_occ_atoms / struct_conn / struct_site Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr1_symmetry / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.ptnr3_symmetry / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr1_symmetry / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_conn.ptnr2_symmetry / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords RNA /
RNA /  APTAMER / RNA-LIGAND COMPLEX /
APTAMER / RNA-LIGAND COMPLEX /  MRNA
MRNA ACETATE ION /
ACETATE ION /  : / COBALT HEXAMMINE(III) /
: / COBALT HEXAMMINE(III) /  RNA / RNA (> 10)
RNA / RNA (> 10) Function and homology information
Function and homology information
 BACILLUS SUBTILIS (bacteria)
BACILLUS SUBTILIS (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.59 Å
MOLECULAR REPLACEMENT / Resolution: 1.59 Å  Authors
Authors Citation
Citation Journal: Chem. Biol. / Year: 2011
Journal: Chem. Biol. / Year: 2011 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2xnz.cif.gz
2xnz.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2xnz.ent.gz
pdb2xnz.ent.gz PDB format
PDB format 2xnz.json.gz
2xnz.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/xn/2xnz
https://data.pdbj.org/pub/pdb/validation_reports/xn/2xnz ftp://data.pdbj.org/pub/pdb/validation_reports/xn/2xnz
ftp://data.pdbj.org/pub/pdb/validation_reports/xn/2xnz


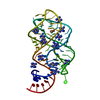
 Links
Links Assembly
Assembly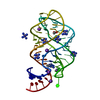
 Components
Components
 BACILLUS SUBTILIS (bacteria)
BACILLUS SUBTILIS (bacteria)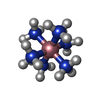

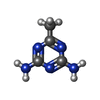






 Acetate
Acetate Acetoguanamine
Acetoguanamine Water
Water X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation
 SYNCHROTRON / Site:
SYNCHROTRON / Site:  ESRF
ESRF  / Beamline: ID14-1 / Wavelength: 0.93344
/ Beamline: ID14-1 / Wavelength: 0.93344  : 0.93344 Å / Relative weight: 1
: 0.93344 Å / Relative weight: 1  Processing
Processing :
:  MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller










 PDBj
PDBj
































