[English] 日本語
 Yorodumi
Yorodumi- PDB-1j87: HUMAN HIGH AFFINITY FC RECEPTOR FC(EPSILON)RI(ALPHA), HEXAGONAL C... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1j87 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
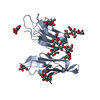
| Title | HUMAN HIGH AFFINITY FC RECEPTOR FC(EPSILON)RI(ALPHA), HEXAGONAL CRYSTAL FORM 1 | |||||||||
 Components Components | HIGH AFFINITY IMMUNOGLOBULIN EPSILON RECEPTOR ALPHA-SUBUNIT | |||||||||
 Keywords Keywords |  IMMUNE SYSTEM / IMMUNE SYSTEM /  Fc Receptor / Fc Receptor /  IgE receptor / IgE receptor /  Glycoprotein Glycoprotein | |||||||||
| Function / homology |  Function and homology information Function and homology informationhigh-affinity IgE receptor activity /  type I hypersensitivity / eosinophil degranulation / IgE binding / type 2 immune response / Fc epsilon receptor (FCERI) signaling / mast cell degranulation / immunoglobulin mediated immune response / Role of LAT2/NTAL/LAB on calcium mobilization / FCERI mediated Ca+2 mobilization ...high-affinity IgE receptor activity / type I hypersensitivity / eosinophil degranulation / IgE binding / type 2 immune response / Fc epsilon receptor (FCERI) signaling / mast cell degranulation / immunoglobulin mediated immune response / Role of LAT2/NTAL/LAB on calcium mobilization / FCERI mediated Ca+2 mobilization ...high-affinity IgE receptor activity /  type I hypersensitivity / eosinophil degranulation / IgE binding / type 2 immune response / Fc epsilon receptor (FCERI) signaling / mast cell degranulation / immunoglobulin mediated immune response / Role of LAT2/NTAL/LAB on calcium mobilization / FCERI mediated Ca+2 mobilization / FCERI mediated MAPK activation / FCERI mediated NF-kB activation / cell surface receptor signaling pathway / external side of plasma membrane / type I hypersensitivity / eosinophil degranulation / IgE binding / type 2 immune response / Fc epsilon receptor (FCERI) signaling / mast cell degranulation / immunoglobulin mediated immune response / Role of LAT2/NTAL/LAB on calcium mobilization / FCERI mediated Ca+2 mobilization / FCERI mediated MAPK activation / FCERI mediated NF-kB activation / cell surface receptor signaling pathway / external side of plasma membrane /  cell surface / cell surface /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 3.2 Å MOLECULAR REPLACEMENT / Resolution: 3.2 Å | |||||||||
 Authors Authors | Garman, S.C. / Sechi, S. / Kinet, J.P. / Jardetzky, T.S. | |||||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2001 Journal: J.Mol.Biol. / Year: 2001Title: The analysis of the human high affinity IgE receptor Fc epsilon Ri alpha from multiple crystal forms. Authors: Garman, S.C. / Sechi, S. / Kinet, J.P. / Jardetzky, T.S. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1j87.cif.gz 1j87.cif.gz | 50.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1j87.ent.gz pdb1j87.ent.gz | 35.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1j87.json.gz 1j87.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/j8/1j87 https://data.pdbj.org/pub/pdb/validation_reports/j8/1j87 ftp://data.pdbj.org/pub/pdb/validation_reports/j8/1j87 ftp://data.pdbj.org/pub/pdb/validation_reports/j8/1j87 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1j86C  1j88C  1j89C  1f2qS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||
| Unit cell |
| ||||||||||
| Details | The biological assembly is a protein monomer with attached carbohydrate |
- Components
Components
| #1: Protein | Mass: 19940.119 Da / Num. of mol.: 1 / Fragment: EXTRACELLULAR FRAGMENT / Mutation: I170C Source method: isolated from a genetically manipulated source Details: GLYCOSYLATED PROTEIN / Source: (gene. exp.)   Homo sapiens (human) / Plasmid: PVL1392 / Production host: Homo sapiens (human) / Plasmid: PVL1392 / Production host:   Spodoptera frugiperda (fall armyworm) / Strain (production host): SF9 / References: UniProt: P12319 Spodoptera frugiperda (fall armyworm) / Strain (production host): SF9 / References: UniProt: P12319 | ||||
|---|---|---|---|---|---|
| #2: Polysaccharide |  / Mass: 424.401 Da / Num. of mol.: 2 / Mass: 424.401 Da / Num. of mol.: 2Source method: isolated from a genetically manipulated source #3: Polysaccharide | alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2- ...alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose |  / Mass: 910.823 Da / Num. of mol.: 1 / Mass: 910.823 Da / Num. of mol.: 1Source method: isolated from a genetically manipulated source #4: Polysaccharide | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta- ...2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose |  / Mass: 570.542 Da / Num. of mol.: 1 / Mass: 570.542 Da / Num. of mol.: 1Source method: isolated from a genetically manipulated source |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.85 Å3/Da / Density % sol: 55 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 5.6 Details: PEG 4000, Sodium Citrate, Sodium Chloride, HECAMEG, pH 5.6, VAPOR DIFFUSION, HANGING DROP at 293K | ||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 23 ℃ / pH: 8.5 / Details: Garman, S.C., (2000) Nature, 406, 259. | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 110 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 5ID-B / Wavelength: 1.004 Å / Beamline: 5ID-B / Wavelength: 1.004 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Nov 15, 1997 |
| Radiation | Monochromator: Si / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.004 Å / Relative weight: 1 : 1.004 Å / Relative weight: 1 |
| Reflection | Resolution: 3.2→20 Å / Num. all: 4076 / Num. obs: 4076 / % possible obs: 92.9 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 8 % / Biso Wilson estimate: 78.1 Å2 / Rmerge(I) obs: 0.103 / Net I/σ(I): 18.3 |
| Reflection shell | Resolution: 3.2→3.31 Å / Redundancy: 7.1 % / Rmerge(I) obs: 0.516 / % possible all: 95.4 |
| Reflection | *PLUS Lowest resolution: 20 Å / % possible obs: 93.2 % / Num. measured all: 32661 |
| Reflection shell | *PLUS % possible obs: 95.4 % / Mean I/σ(I) obs: 5.3 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1F2Q Resolution: 3.2→19.23 Å / Rfactor Rfree error: 0.015 / Data cutoff high absF: 1638969.31 / Data cutoff low absF: 0 / Isotropic thermal model: GROUP / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Engh & Huber Details: no density appears for residues 33-35; they are not included in the model
| |||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 13.72 Å2 / ksol: 0.25 e/Å3 | |||||||||||||||||||||||||
| Displacement parameters | Biso mean: 96 Å2
| |||||||||||||||||||||||||
| Refine analyze |
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3.2→19.23 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 3.2→3.4 Å / Rfactor Rfree error: 0.049 / Total num. of bins used: 6
| |||||||||||||||||||||||||
| Xplor file |
| |||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 1 / Classification: refinement | |||||||||||||||||||||||||
| Refinement | *PLUS σ(F): 0 / % reflection Rfree: 10.2 % | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 96 Å2 | |||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| |||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.393 / % reflection Rfree: 10 % / Rfactor Rwork: 0.32 |
 Movie
Movie Controller
Controller












 PDBj
PDBj








