+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 1c38 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | SOLUTION STRUCTURE OF A QUADRUPLEX FORMING DNA AND ITS INTERMEDIATE | ||||||||||||||||||
 要素 要素 | DNA (5'-D(* キーワード キーワード DNA (デオキシリボ核酸) / DNA (デオキシリボ核酸) /  APTAMER (アプタマー) / APTAMER (アプタマー) /  THROMBIN BINDING (トロンビン) / THROMBIN BINDING (トロンビン) /  POTASSIUM (カリウム) / METAL ION BINDING SITES / QUADRUPLEX / INTERMEDIATE POTASSIUM (カリウム) / METAL ION BINDING SITES / QUADRUPLEX / INTERMEDIATE機能・相同性 | : / |  デオキシリボ核酸 / DNA (> 10) デオキシリボ核酸 / DNA (> 10) 機能・相同性情報 機能・相同性情報手法 |  溶液NMR / THE DNA-POTASSIUM STRUCTURE WAS MINIMIZED IN 500 STEPS OF POWELL'S CONJUGATE GRADIENT MINIMIZATION. 溶液NMR / THE DNA-POTASSIUM STRUCTURE WAS MINIMIZED IN 500 STEPS OF POWELL'S CONJUGATE GRADIENT MINIMIZATION.  データ登録者 データ登録者Bolton, P.H. / Marathias, V.M. / Wang, K. |  引用 引用 ジャーナル: Nucleic Acids Res. / 年: 2000 ジャーナル: Nucleic Acids Res. / 年: 2000タイトル: Structures of the potassium-saturated, 2:1, and intermediate, 1:1, forms of a quadruplex DNA. 著者: Marathias, V.M. / Bolton, P.H. #1:  ジャーナル: J.Mol.Biol. / 年: 1996 ジャーナル: J.Mol.Biol. / 年: 1996タイトル: Determination of the Number and Location of the Manganese Binding Sites of DNA Quadruplexes in Solution by EPR and NMR in the Presence and Absence of Thrombin 著者: Bolton, P.H. / Marathias, V.M. / Wang, K. 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  1c38.cif.gz 1c38.cif.gz | 31.7 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb1c38.ent.gz pdb1c38.ent.gz | 17.4 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  1c38.json.gz 1c38.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/c3/1c38 https://data.pdbj.org/pub/pdb/validation_reports/c3/1c38 ftp://data.pdbj.org/pub/pdb/validation_reports/c3/1c38 ftp://data.pdbj.org/pub/pdb/validation_reports/c3/1c38 | HTTPS FTP |
|---|
-関連構造データ
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 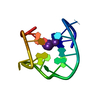
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR アンサンブル |
|
- 要素
要素
| #1: DNA鎖 | 分子量: 4743.051 Da / 分子数: 1 / 由来タイプ: 合成 詳細: THROMBIN BINDING APTAMER INTERMEDIATE WITH MULTIPLE POTASSIUM MINIMIZATIONS |
|---|---|
| #2: 化合物 | ChemComp-K / |
-実験情報
-実験
| 実験 | 手法:  溶液NMR 溶液NMR |
|---|---|
| NMR実験の詳細 | Text: THE 10 POTASSIUM POSITIONS IN EACH QUARTET ARE THE LOWEST ENERGY POSITION CALCULATED USING X-PLOR AND BASED ON THE REFINED SOLUTION STRUCTURE OF THE INTERMEDIATE APTAMER QUADRUPLEX. THE ...Text: THE 10 POTASSIUM POSITIONS IN EACH QUARTET ARE THE LOWEST ENERGY POSITION CALCULATED USING X-PLOR AND BASED ON THE REFINED SOLUTION STRUCTURE OF THE INTERMEDIATE APTAMER QUADRUPLEX. THE DISPERSION OF THE POTASSIUM POSITION INDICATE THE BINDING SITE STABILITY. |
- 試料調製
試料調製
結晶化 | *PLUS 手法: その他 / 詳細: NMR |
|---|
- 解析
解析
| NMR software | 名称:  X-PLOR / バージョン: 3.1 / 開発者: BRUNGER / 分類: 精密化 X-PLOR / バージョン: 3.1 / 開発者: BRUNGER / 分類: 精密化 |
|---|---|
| 精密化 | 手法: THE DNA-POTASSIUM STRUCTURE WAS MINIMIZED IN 500 STEPS OF POWELL'S CONJUGATE GRADIENT MINIMIZATION. ソフトェア番号: 1 詳細: X-PLOR 3.1 WAS USED TO DETERMINE THE MINIMUM ENERGY POSITIONS OF THE POTASSIUM IONS. A VAN DER WALLS RADIUS OF 1.96 A AND A CHARGE OF PLUS ONE WAS USED FOR EACH POTASSIUM ION. A DISTANCE- ...詳細: X-PLOR 3.1 WAS USED TO DETERMINE THE MINIMUM ENERGY POSITIONS OF THE POTASSIUM IONS. A VAN DER WALLS RADIUS OF 1.96 A AND A CHARGE OF PLUS ONE WAS USED FOR EACH POTASSIUM ION. A DISTANCE-DEPENDENT DIELECTRIC FUNCTION WAS INCLUDED IN THE NON-BONDED TERMS. THE OPTIONS FOR THE DISTANCE-DEPENDENT DIELECTRIC FUNCTION USED IN THIS PROTOCOL INVOLVE USING A VAN DER WAALS FUNCTION IN COMBINATION WITH A ELECTROSTATIC FUNCTION AND A 1/R-DEPENDENT DIELECTRIC FUNCTION, WHERE R INDICATES THE DISTANCE BETWEEN ATOMS (COULOMB'S LAW). IN ORDER TO SIMULATE AN AQUEOUS ENVIRONMENT, THE DIELECTRIC CONSTANT OF THE SYSTEM WAS SET TO 64. THE SWITCHING FUNCTION FOR THE NON-BONDED INTERACTIONS WAS ACTIVE BETWEEN 7.5 AND 10.0 A. THE NON-BONDED INTERACTION CUT-OFF WAS SET TO BE A SMOOTH TRANSITION BETWEEN 0.5 AND 10.0 A. THE DNA/POTASSIUM COMPLEXES WERE SUBJECTED TO A 500-STEP MINIMIZATION ALLOWING ONLY THE POTASSIUM TO MOVE WHILE THE DNA WAS HELD RIGID. ENERGY MINIMIZATION'S WERE RUN WITH VARIOUS STARTING POSITIONS FOR THE POTASSIUM IONS IN ORDER TO DETERMINE IF THE INITIAL POSITION HAD AN EFFECT ON THE MINIMIZED POSITION. RANDOM STARTING POSITIONS OF THE K+ FROM 5 TO 10 A AWAY FROM THE DNA WERE USED AND THE FINAL RESULTS WERE FOUND TO BE INDEPENDENT OF THE INITIAL COORDINATES OF THE POTASSIUM. THE MINIMIZATIONS WERE CARRIED OUT FIRST WITH ONE POTASSIUM ION PER QUADRUPLEX AND THEN WITH TWO PER QUADRUPLEX. THE POSITION OF THE FIRST BINDING SITE DID NOT CHANGE, UPON ADDITION OF THE SECOND POTASSIUM, FOR EITHER DNA. ADDITIONAL MINIMIZATIONS DID NOT CHANGE THE POSITIONS OF THE POTASSIUM IONS. |
| NMRアンサンブル | 登録したコンフォーマーの数: 1 |
 ムービー
ムービー コントローラー
コントローラー









 PDBj
PDBj








































