+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram average of membrane-bound Atg18 oligomers | |||||||||
 Map data Map data | sharp map from Relion Software | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  autophagy / membrane remodeling / PIP binding / autophagy / membrane remodeling / PIP binding /  PI3P / PI(3 / 5)P2 / lipid binding protein / PI3P / PI(3 / 5)P2 / lipid binding protein /  MEMBRANE PROTEIN MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of phosphatidylinositol biosynthetic process / PAS complex / 1-phosphatidyl-1D-myo-inositol 3,5-bisphosphate metabolic process / phagophore /  vacuolar protein processing / positive regulation of vacuole organization / vacuolar protein processing / positive regulation of vacuole organization /  Macroautophagy / cytoplasm to vacuole targeting by the Cvt pathway / nucleophagy / pexophagy ...regulation of phosphatidylinositol biosynthetic process / PAS complex / 1-phosphatidyl-1D-myo-inositol 3,5-bisphosphate metabolic process / phagophore / Macroautophagy / cytoplasm to vacuole targeting by the Cvt pathway / nucleophagy / pexophagy ...regulation of phosphatidylinositol biosynthetic process / PAS complex / 1-phosphatidyl-1D-myo-inositol 3,5-bisphosphate metabolic process / phagophore /  vacuolar protein processing / positive regulation of vacuole organization / vacuolar protein processing / positive regulation of vacuole organization /  Macroautophagy / cytoplasm to vacuole targeting by the Cvt pathway / nucleophagy / pexophagy / protein localization to phagophore assembly site / piecemeal microautophagy of the nucleus / phagophore assembly site membrane / late endosome to vacuole transport / Macroautophagy / cytoplasm to vacuole targeting by the Cvt pathway / nucleophagy / pexophagy / protein localization to phagophore assembly site / piecemeal microautophagy of the nucleus / phagophore assembly site membrane / late endosome to vacuole transport /  phosphatidylinositol-3-phosphate binding / fungal-type vacuole membrane / phagophore assembly site / phosphatidylinositol-4-phosphate binding / phosphatidylinositol-3,5-bisphosphate binding / vacuolar membrane / phosphatidylinositol-3-phosphate binding / fungal-type vacuole membrane / phagophore assembly site / phosphatidylinositol-4-phosphate binding / phosphatidylinositol-3,5-bisphosphate binding / vacuolar membrane /  extrinsic component of membrane / extrinsic component of membrane /  autophagosome assembly / autophagosome assembly /  ubiquitin binding / cell periphery / ubiquitin binding / cell periphery /  macroautophagy / macroautophagy /  protein transport / endosome membrane / protein transport / endosome membrane /  endosome / protein-containing complex / endosome / protein-containing complex /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   Saccharomyces cerevisiae (brewer's yeast) Saccharomyces cerevisiae (brewer's yeast) | |||||||||
| Method | subtomogram averaging /  cryo EM / Resolution: 26.0 Å cryo EM / Resolution: 26.0 Å | |||||||||
 Authors Authors | Mann D / Fromm S / Martinez-Sanchez A / Gopaldass N / Mayer A / Sachse C | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structures of Atg18 oligomers reveal a tilted structural scaffold for Atg2 at the isolation membrane Authors: Mann D / Fromm SA / Martinez-Sanchez A / Gopaldass N / Mayer A / Sachse C | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15412.map.gz emd_15412.map.gz | 22 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15412-v30.xml emd-15412-v30.xml emd-15412.xml emd-15412.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15412.png emd_15412.png | 30.4 KB | ||
| Filedesc metadata |  emd-15412.cif.gz emd-15412.cif.gz | 5.7 KB | ||
| Others |  emd_15412_half_map_1.map.gz emd_15412_half_map_1.map.gz emd_15412_half_map_2.map.gz emd_15412_half_map_2.map.gz | 22 MB 22 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15412 http://ftp.pdbj.org/pub/emdb/structures/EMD-15412 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15412 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15412 | HTTPS FTP |
-Related structure data
| Related structure data |  8afyMC  8afqC  8afwC  8afxC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15412.map.gz / Format: CCP4 / Size: 28.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15412.map.gz / Format: CCP4 / Size: 28.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharp map from Relion Software | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.129 Å | ||||||||||||||||||||||||||||||||||||
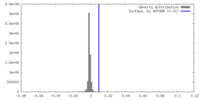
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map B
| File | emd_15412_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
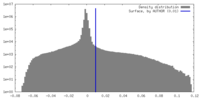
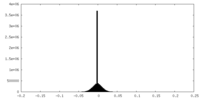
| Density Histograms |
-Half map: half map A
| File | emd_15412_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
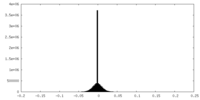
| Density Histograms |
- Sample components
Sample components
-Entire : Subtomogram average of membrane-bound Atg18
| Entire | Name: Subtomogram average of membrane-bound Atg18 |
|---|---|
| Components |
|
-Supramolecule #1: Subtomogram average of membrane-bound Atg18
| Supramolecule | Name: Subtomogram average of membrane-bound Atg18 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Saccharomyces cerevisiae (brewer's yeast) Saccharomyces cerevisiae (brewer's yeast) |
-Macromolecule #1: Autophagy-related protein 18
| Macromolecule | Name: Autophagy-related protein 18 / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Saccharomyces cerevisiae (brewer's yeast) / Strain: ATCC 204508 / S288c Saccharomyces cerevisiae (brewer's yeast) / Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 55.045777 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MSDSSPTINF INFNQTGTCI SLGTSKGFKI FNCEPFGKFY SEDSGGYAIV EMLFSTSLLA LVGIGDQPAL SAARLRIINT KKHSIICEV TFPTSILSVK MNKSRLVVLL QEQIYIYDIN TMRLLHTIET NPNPRGLMAM SPSVANSYLV YPSPPKVINS E IKAHATTN ...String: MSDSSPTINF INFNQTGTCI SLGTSKGFKI FNCEPFGKFY SEDSGGYAIV EMLFSTSLLA LVGIGDQPAL SAARLRIINT KKHSIICEV TFPTSILSVK MNKSRLVVLL QEQIYIYDIN TMRLLHTIET NPNPRGLMAM SPSVANSYLV YPSPPKVINS E IKAHATTN NITLSVGGNT ETSFKRDQQD AGHSDISDLD QYSSFTKRDD ADPTSSNGGN SSIIKNGDVI VFNLETLQPT MV IEAHKGE IAAMAISFDG TLMATASDKG TIIRVFDIET GDKIYQFRRG TYATRIYSIS FSEDSQYLAV TGSSKTVHIF KLG HSMSNN KLDSDDSNME EAAADDSSLD TTSIDALSDE ENPTRLAREP YVDASRKTMG RMIRYSSQKL SRRAARTLGQ IFPI KVTSL LESSRHFASL KLPVETNSHV MTISSIGSPI DIDTSEYPEL FETGNSASTE SYHEPVMKMV PIRVVSSDGY LYNFV MDPE RGGDCLILSQ YSILMD UniProtKB: Autophagy-related protein 18 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 99 % / Chamber temperature: 291 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 0.0 µm / Nominal defocus min: 0.0 µm Bright-field microscopy / Nominal defocus max: 0.0 µm / Nominal defocus min: 0.0 µm |
| Specialist optics | Phase plate: VOLTA PHASE PLATE |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 3.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Extraction | Number tomograms: 16 / Number images used: 8300 |
|---|---|
| Final 3D classification | Software - Name: RELION |
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: RELION |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 26.0 Å / Resolution method: OTHER / Software - Name: RELION / Details: Mask-less FDR-FSC at 1% FDR / Number subtomograms used: 8300 |
-Atomic model buiding 1
| Details | Fitted 4x dimer "T" from Atg18 filament structure to interpret the density as a tetramer of dimers |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-8afy: |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)