-検索条件
-検索結果
検索 (著者・登録者: mina & p)の結果248件中、1から50件目までを表示しています

EMDB-36659: 
Structure of human TRPV4 with antagonist A1

EMDB-36660: 
Structure of human TRPV4 with antagonist GSK279
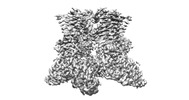
EMDB-36675: 
Structure of human TRPV4 with antagonist A2

EMDB-36676: 
Structure of human TRPV4 with antagonist A2 and RhoA

PDB-8ju5: 
Structure of human TRPV4 with antagonist A1

PDB-8ju6: 
Structure of human TRPV4 with antagonist GSK279

PDB-8jvi: 
Structure of human TRPV4 with antagonist A2

PDB-8jvj: 
Structure of human TRPV4 with antagonist A2 and RhoA

EMDB-42845: 
Selenocysteine synthase- SelA

PDB-8uzw: 
Selenocysteine synthase- SelA

EMDB-18659: 
Cryo-EM Structure of Human Kv3.1 in Complex with Modulator AUT1

EMDB-18660: 
Cryo-EM Structure of Human Kv3.1 in Complex with Modulator AUT5

PDB-8quc: 
Cryo-EM Structure of Human Kv3.1 in Complex with Modulator AUT1

PDB-8qud: 
Cryo-EM Structure of Human Kv3.1 in Complex with Modulator AUT5

EMDB-37465: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by sucrose density

EMDB-37466: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by Ca2+-DEAE

PDB-8wdu: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by sucrose density

PDB-8wdv: 
Photosynthetic LH1-RC complex from the purple sulfur bacterium Allochromatium vinosum purified by Ca2+-DEAE
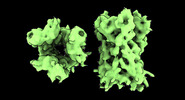
EMDB-17704: 
Subtomogram average of Vaccinia A10 trimer with open center from in vitro cores
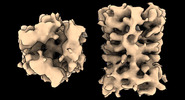
EMDB-17708: 
Subtomogram average of Vaccinia A10 trimer with tight center from in vitro cores

EMDB-17753: 
Subtomogram average of Vaccinia A10 trimer from in situ cores

EMDB-43013: 
Myxococcus xanthus HEnc-K417N(A) protein shell with icosahedral T=1 symmetry

EMDB-43016: 
Myxococcus xanthus HEnc-K417N(A) protein shell with tetrahedral symmetry (12 pentamers, 4 hexamers)

EMDB-43037: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D3 symmetry (12 pentamers, 3 hexamers)

EMDB-43038: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D6 symmetry (12 pentamers, 8 hexamers)

EMDB-43039: 
Myxococcus xanthus HEnc-K417N(A) protein shell with C2 symmetry (12 pentamers, 9 hexamers)

EMDB-43040: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D3 symmetry (12 pentamers, 11 hexamers)

EMDB-43041: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D2 symmetry (12 pentamers, 12 hexamers)

EMDB-43042: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D2 symmetry (12 pentamers, 14 hexamers)

EMDB-43043: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D5 symmetry (12 pentamers, 15 hexamers)

EMDB-43113: 
Myxococcus xanthus EncA WT protein shell with icosahedral symmetry T=3

EMDB-15359: 
Structure of dimeric yeast RNA polymerase II bound to a transcription bubble (focused map of monomer 1)

EMDB-15360: 
Structure of dimeric yeast RNA polymerase II bound to a transcription bubble (focused map of monomer 2)

EMDB-41617: 
CryoEM structure of PI3Kalpha

PDB-8tu6: 
CryoEM structure of PI3Kalpha

EMDB-15358: 
Structure of dimeric yeast RNA polymerase II bound to a transcription bubble (consensus map)

EMDB-27703: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace

PDB-8dtk: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace

EMDB-29930: 
T. cruzi topoisomerase II alpha bound to dsDNA and the covalent inhibitor CT1

PDB-8gcc: 
T. cruzi topoisomerase II alpha bound to dsDNA and the covalent inhibitor CT1

EMDB-16325: 
Cryo-EM structure of SKP1-SKP2-CKS1-CDK2-CyclinA-p27KIP1 Complex
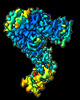
EMDB-16327: 
Cryo-EM structure of SKP1-SKP2-CKS1 from the SCFSKP2 E3 ligase complex
EMDB-16344: 
Cryo-EM structure of CDK2-CyclinA in complex with p27 from the SCFSKP2 E3 ligase Complex

PDB-8bya: 
Cryo-EM structure of SKP1-SKP2-CKS1-CDK2-CyclinA-p27KIP1 Complex

PDB-8byl: 
Cryo-EM structure of SKP1-SKP2-CKS1 from the SCFSKP2 E3 ligase complex

PDB-8bzo: 
Cryo-EM structure of CDK2-CyclinA in complex with p27 from the SCFSKP2 E3 ligase Complex

EMDB-26735: 
Hantavirus ANDV Gn(H) protein in complex with 2 Fabs ANDV-5 and ANDV-34
EMDB-26736: 
Hantavirus MAPV Gn(H)/Gc protein in complex with 2 Fabs SNV-24 and SNV-53

EMDB-27318: 
CryoEM structure of Hantavirus ANDV Gn(H) protein complex with 2Fabs ANDV-5 and ANDV-34

PDB-8dbz: 
CryoEM structure of Hantavirus ANDV Gn(H) protein complex with 2Fabs ANDV-5 and ANDV-34
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
