-Search query
-Search result
Showing all 47 items for (author: schwab & ra)

EMDB-16128: 
Tomogram of a late endosome of A549 cell infected with influenza A virus.

EMDB-16129: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6C).

EMDB-16130: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6E)

EMDB-16131: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6G)

EMDB-16132: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6I,K,M)

EMDB-16133: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6O)

EMDB-15705: 
Tomogram of a late endosome of A549 cell (Figure 1R)

EMDB-15707: 
Tomogram of a late endosome of A549 cell treated with IFN-beta (Figure 1T)
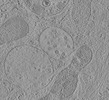
EMDB-15708: 
Tomogram of a late endosome of A549-IFITM3 cell (Figure 1V)

EMDB-15131: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S7)
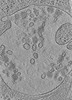
EMDB-15130: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure 4E)
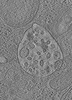
EMDB-15132: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S8)

EMDB-15133: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S9)

EMDB-13764: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA (Composite Map)

PDB-7q1u: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA (Composite Map)

EMDB-14578: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA (Consensus Map)

EMDB-13860: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to IMP-1575

PDB-7q6z: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to IMP-1575

EMDB-13841: 
Focused refinement of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA

EMDB-13842: 
Focused refinement of Hedgehog acyltransferase (HHAT) in complex with megabody 177 - megabody core

EMDB-12940: 
SARS-CoV-2-Induced Reshaping of Subcellular Morphologies

EMDB-12286: 
Cryo-EM structure of the ternary complex between Netrin-1, Neogenin and Repulsive Guidance Molecule B

PDB-7ndg: 
Cryo-EM structure of the ternary complex between Netrin-1, Neogenin and Repulsive Guidance Molecule B

EMDB-11938: 
Ternary complex of full-length Caspase-8 and FADD

EMDB-11939: 
Central region of Caspase-8:FADD ternary complex

EMDB-11940: 
Ternary complex of Full length Caspase-8 with FADD and FLIPs

EMDB-11941: 
CryoEM analysis of ternary complex of full-length Caspase-8 with FADD and FLIPs

EMDB-11837: 
Structure of the MTA1/HDAC1/MBD2 NURD deacetylase complex

EMDB-11838: 
Structure of the core MTA1/HDAC1/MBD2 NURD deacetylase complex

EMDB-11839: 
Structure of the extended MTA1/HDAC1/MBD2/RBBP4 NURD deacetylase complex

PDB-7ao8: 
Structure of the MTA1/HDAC1/MBD2 NURD deacetylase complex

PDB-7ao9: 
Structure of the core MTA1/HDAC1/MBD2 NURD deacetylase complex

PDB-7aoa: 
Structure of the extended MTA1/HDAC1/MBD2/RBBP4 NURD deacetylase complex

EMDB-10660: 
Nup116_delta_NPC_25C

EMDB-10661: 
Nup116delta_NPC_37C

EMDB-10198: 
In-cell S cerevisiae nuclear pore complex

EMDB-11041: 
The structure of the dimeric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex

EMDB-11042: 
The structure of the tetrameric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex

PDB-6z2j: 
The structure of the dimeric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex

PDB-6z2k: 
The structure of the tetrameric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex

EMDB-10626: 
Negative stain map of CoREST complex (LSD1:RCOR1:HDAC1)

EMDB-10627: 
Cryo-EM map of glutaraldehye cross-linked CoREST complex (LSD1:RCOR1:HDAC1)

EMDB-10628: 
Cryo EM map of BS3 crosslinked CoREST complex (LSD1:RCOR1:HDAC1)- closed form

EMDB-10629: 
Cryo EM map of BS3 crosslinked CoREST complex (LSD1:RCOR1:HDAC1) - open form

EMDB-10630: 
Interaction of the CoREST complex with a nucleosome with 185 bp 601 sequence DNA and a propargylamine mimic of dimethy Lys4 histone H3

PDB-6rvd: 
Revised cryo-EM structure of the human 2:1 Ptch1-Shh complex

EMDB-3399: 
Structure of the core NuRD complex (MTA1:HDAC1:RBBP4)
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
