-Search query
-Search result
Showing 1 - 50 of 135 items for (author: nans & a)

EMDB-17449: 
S. cerevisiae nexus-sCMGE after DNA replication initiation

EMDB-17458: 
S. cerevisiae ssDNA-sCMGE after DNA replication initiation

EMDB-17459: 
S. cerevisiae consensus-sCMGE on ssDNA after DNA replication initiation

EMDB-17460: 
S. cerevisiae sCMGE with N-ter Mcm10 density

PDB-8p5e: 
S. cerevisiae nexus-sCMGE after DNA replication initiation

PDB-8p62: 
S. cerevisiae ssDNA-sCMGE after DNA replication initiation

PDB-8p63: 
S. cerevisiae consensus-sCMGE on ssDNA after DNA replication initiation

EMDB-18729: 
Cryo-EM structure of tetrameric human SAMHD1 with dApNHpp

EMDB-18730: 
Cryo-EM structure of tetrameric human SAMHD1 State I - Tense

EMDB-18731: 
Cryo-EM structure of tetrameric human SAMHD1 State II - Hemi-relaxed

EMDB-18732: 
Cryo-EM structure of tetrameric human SAMHD1 State III - Relaxed

EMDB-18733: 
Cryo-EM structure of tetrameric human SAMHD1 State IV - Depleted relaxed

EMDB-18734: 
Cryo-EM structure of tetrameric human SAMHD1 State V - Depleted relaxed

PDB-8qxj: 
Cryo-EM structure of tetrameric human SAMHD1 with dApNHpp

PDB-8qxk: 
Cryo-EM structure of tetrameric human SAMHD1 State I - Tense

PDB-8qxl: 
Cryo-EM structure of tetrameric human SAMHD1 State II - Hemi-relaxed

PDB-8qxm: 
Cryo-EM structure of tetrameric human SAMHD1 State III - Relaxed

PDB-8qxn: 
Cryo-EM structure of tetrameric human SAMHD1 State IV - Depleted relaxed

PDB-8qxo: 
Cryo-EM structure of tetrameric human SAMHD1 State V - Depleted relaxed
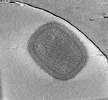
EMDB-18916: 
Cryotomogram of mature Vaccinia virus (WR) virion

EMDB-18917: 
Subtomogram average of the Vaccinia virus (WR) portal complex in mature virions

EMDB-18918: 
Subtomogram average of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions

PDB-8r5i: 
In situ structure of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions by flexible fitting into a cryoET map

EMDB-15596: 
In situ subtomogram average of Vaccinia virus (WR) palisade, all virions

EMDB-15597: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from CEVs

EMDB-15598: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from IMVs

EMDB-15599: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from IEVs

EMDB-15600: 
Cryotomogram of Vaccinia virus (WR) infected HeLa cell (immature virions)

EMDB-15601: 
Cryotomogram of Vaccinia virus (WR) infected HeLa cell (mature virions)

EMDB-15602: 
In situ subtomogram average of Vaccinia virus (WR) D13 lattice, on immature virions

PDB-8arh: 
In situ subtomogram average of Vaccinia virus (WR) D13 lattice, on immature virions

EMDB-26862: 
CryoEM structure of the TIR domain from AbTir in complex with 3AD

PDB-7uxu: 
CryoEM structure of the TIR domain from AbTir in complex with 3AD

EMDB-13978: 
S. cerevisiae CMGE nucleating origin DNA melting

EMDB-13988: 
S. cerevisiae CMGE dimer nucleating origin DNA melting

EMDB-14439: 
S. cerevisiae CMGE dimer nucleating origin DNA melting

PDB-7qhs: 
S. cerevisiae CMGE nucleating origin DNA melting

PDB-7z13: 
S. cerevisiae CMGE dimer nucleating origin DNA melting

EMDB-13439: 
Cryo-EM structure of E. coli TnsB in complex with right end fragment of Tn7 transposon

EMDB-13440: 
Cryo-EM helical reconstruction of E. coli TnsB in complex with right end fragment of Tn7 transposon

EMDB-26322: 
MVV cleaved synaptic complex (CSC) intasome at 3.4 A resolution

PDB-7u32: 
MVV cleaved synaptic complex (CSC) intasome at 3.4 A resolution

EMDB-26191: 
Cryo-EM structure of human SARM1 TIR domain in complex with 1AD

EMDB-14453: 
MVV strand transfer complex (STC) intasome in complex with LEDGF/p75 at 3.5 A resolution

EMDB-24272: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (TIR:1AD)

EMDB-24273: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (ARM and SAM domains)

EMDB-24274: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (SAM-TIR:1AD)

PDB-7nak: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (TIR:1AD)

PDB-7nal: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (ARM and SAM domains)

EMDB-13176: 
3.0 A resolution structure of a DNA-loaded MCM double hexamer
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
