-Search query
-Search result
Showing all 47 items for (author: yassin & as)

EMDB-28092: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-093
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28090: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-040
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28091: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-045
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28093: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-156
Method: single particle / : Shek J, Callaway H, Li H, Yu X, Saphire EO

EMDB-28094: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-234
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28095: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-260
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28096: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-279
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28097: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-290
Method: single particle / : Yu X, Callaway H, Li H, Shek J, Saphire EO
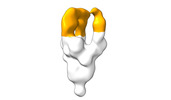
EMDB-28098: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-294
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28099: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-295
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28100: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-299
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28102: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-334
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
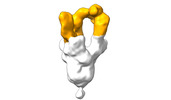
EMDB-28103: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-360
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28104: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-361
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28105: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-362
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28106: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-368
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28168: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-292
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28169: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-333
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
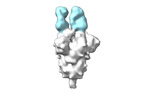
EMDB-28170: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-355
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28171: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-371
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-20911: 
Group 1 H1 influenza stem nanoparticle with a group 2-like N-linked glycosylation site at position 38
Method: single particle / : Tsybovsky Y, Stephens T, Kanekiyo M, Graham BS

EMDB-20912: 
Group 1 H1 influenza stem nanoparticle with an H38R mutation
Method: single particle / : Tsybovsky Y, Stephens T, Kanekiyo M, Graham BS

EMDB-20913: 
Group 1 H1 influenza stem nanoparticle
Method: single particle / : Tsybovsky Y, Stephens T, Kanekiyo M, Graham BS

EMDB-0264: 
Type VI membrane complex
Method: single particle / : Rapisarda C, Fronzes R

EMDB-0265: 
Type VI membrane complex
Method: single particle / : Rapisarda C, Fronzes R

EMDB-0266: 
Type VI membrane complex
Method: single particle / : Rapisarda C, Fronzes R

EMDB-0267: 
Type VI secretion membrane complex
Method: single particle / : Rapisarda C, Fronzes R

EMDB-4562: 
Subtomogram average of T6SS membrane complex TssJLM
Method: subtomogram averaging / : Kooger R, Pilhofer M

PDB-6hs7: 
Type VI membrane complex
Method: single particle / : Rapisarda C, Fronzes R

EMDB-7021: 
Structure of influenza hemagglutinin from A/New Caledonia/20/99 in complex with FAB 441D6.
Method: single particle / : Gallagher JR, Harris AK

EMDB-0008: 
The baseplate complex from the type VI secretion system
Method: single particle / : Rapisarda C, Fronzes R

EMDB-0009: 
The baseplate complex from the type VI secretion system
Method: single particle / : Rapisarda C, Fronzes R

EMDB-0010: 
The baseplate complex from the type VI secretion system
Method: single particle / : Rapisarda C, Fronzes R

PDB-6giy: 
The baseplate complex from the type VI secretion system
Method: single particle / : Rapisarda C, Fronzes R

PDB-6gj1: 
The baseplate complex from the type VI secretion system
Method: single particle / : Rapisarda C, Fronzes R

PDB-6gj3: 
The baseplate complex from the type VI secretion system
Method: single particle / : Rapisarda C, Fronzes R

EMDB-8066: 
HKU1 spike with attached foldon domain and wild-type furin-cleavage site
Method: single particle / : Kirchdoerfer RN, Cottrell CA, Wang N, Pallesen J, Yassine HM, Turner HL, Corbett KS, Graham BS, McLellan JS, Ward AB

EMDB-8067: 
HKU1 spike with attached foldon domain and wild-type furin-cleavage site
Method: single particle / : Kirchdoerfer RN, Cottrell CA, Wang N, Pallesen J, Yassine HM, Turner HL, Corbett KS, Graham BS, McLellan JS, Ward AB
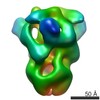
EMDB-8068: 
HKU1 spike with attached foldon domain and wild-type furin-cleavage site
Method: single particle / : Kirchdoerfer RN, Cottrell CA, Wang N, Pallesen J, Yassine HM, Turner HL, Corbett KS, Graham BS, McLellan JS, Ward AB

EMDB-8069: 
Prefusion structure of a human coronavirus spike protein
Method: single particle / : Kirchdoerfer RN, Cottrell CA, Wang N, Pallesen J, Yassine HM, Turner HL, Corbett KS, Graham BS, McLellan JS, Ward AB

PDB-5i08: 
Prefusion structure of a human coronavirus spike protein
Method: single particle / : Kirchdoerfer RN, Cottrell CA, Wang N, Pallesen J, Yassine HM, Turner HL, Corbett KS, Graham BS, McLellan JS, Ward AB

EMDB-6332: 
3D reconstruction of a ferritin-based nanoparticle displaying H1 Hemagglutinin stem epitopes
Method: single particle / : Gallagher JR, Harris AK, Yassine HM, Boyington JC, McTamney PM, Wei CJ, Masaru M, Kong WP, Wang L, Zhang Y, Joyce MG, Lingwood D, Moin SM, Andersen H, Okuno Y, Rao SS, Kwong PD, Mascola JR, Nabel GJ, Graham BS

PDB-3izz: 
Models for ribosome components that are nearest neighbors to the bovine mitochondrial initiation factor2 bound to the E. Coli ribosome
Method: single particle / : Yassin AS, Haque E, Datta PP, Elmore K, Banavali NK, Spremulli LL, Agrawal RK

PDB-3izy: 
Mammalian mitochondrial translation initiation factor 2
Method: single particle / : Yassin AS, Haque E, Datta PP, Elmore K, Banavali NK, Spremulli LL, Agrawal RK

EMDB-1854: 
An insertion domain within mammalian mitochondrial translation initiation factor 2 serves the role of eubacterial initiation factor 1
Method: single particle / : Yassin AS, Haque E, Datta PP, Elmore K, Banavali NK, Spremulli LL, Agrawal RK
EMDB-1855: 
An insertion domain within mammalian mitochondrial translation initiation factor 2 serves the role of eubacterial initiation factor 1
Method: single particle / : Yassin AS, Haque E, Datta PP, Elmore K, Banavali NK, Spremulli LL, Agrawal RK
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
