-Search query
-Search result
Showing 1 - 50 of 247 items for (author: villa & n)

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E
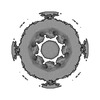
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-40674: 
Subtomogram average of immature PhiKZ Major Capsid Protein from tubular arrays
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-17189: 
Structure of the human neutral amino acid transporter ASCT2 in complex with nanobody 469
Method: single particle / : Canul-Tec J, Reyes N

EMDB-17192: 
Complex of human ASCT2 with Syncytin-1
Method: single particle / : Khare S, Reyes N

EMDB-17193: 
Complex of ASCT2 with Suppressyn
Method: single particle / : Khare S, Kumar A, Reyes N
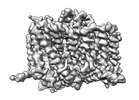
EMDB-17194: 
Heterotrimeric Complex of Human ASCT2 with Syncytin-1
Method: single particle / : Khare S, Reyes N

PDB-8oud: 
Structure of the human neutral amino acid transporter ASCT2 in complex with nanobody 469
Method: single particle / : Canul-Tec J, Reyes N

PDB-8ouj: 
Heterotrimeric Complex of Human ASCT2 with Syncytin-1
Method: single particle / : Khare S, Reyes N

EMDB-17125: 
Knockout of GMC-oxidoreductase genes reveals that functional redundancy preserves mimivirus essential functions
Method: helical / : Alempic JM, Bisio H, Villalta A, Santini S, Lartigue A, Schmitt A, Bugnot C, Notaro A, Belmudes L, Adrait A, Poirot O, Ptchelkine D, De Castro C, Coute Y, Abergel C

EMDB-17131: 
Knockout of GMC-oxidoreductase genes reveals that functional redundancy preserves mimivirus essential functions
Method: helical / : Alempic JM, Bisio H, Villalta A, Santini S, Lartigue A, Schmitt A, Bugnot C, Notaro A, Belmudes L, Adrait A, Poirot O, Ptchelkine D, De Castro C, Coute Y, Abergel C

PDB-8orh: 
Knockout of GMC-oxidoreductase genes reveals that functional redundancy preserves mimivirus essential functions
Method: helical / : Alempic JM, Bisio H, Villalta A, Santini S, Lartigue A, Schmitt A, Bugnot C, Notaro A, Belmudes L, Adrait A, Poirot O, Ptchelkine D, De Castro C, Coute Y, Abergel C

PDB-8ors: 
Knockout of GMC-oxidoreductase genes reveals that functional redundancy preserves mimivirus essential functions
Method: helical / : Alempic JM, Bisio H, Villalta A, Santini S, Lartigue A, Schmitt A, Bugnot C, Notaro A, Belmudes L, Adrait A, Poirot O, Ptchelkine D, De Castro C, Coute Y, Abergel C

EMDB-42455: 
Candidatus Methanomethylophilus alvus tRNAPyl in A-site of ribosome
Method: single particle / : Krahn N, Zhang J, Melnikov SV, Tharp JM, Villa A, Patel A, Howard RJ, Gabir H, Patel TR, Stetefeld J, Puglisi J, Soll D

PDB-8upt: 
Candidatus Methanomethylophilus alvus tRNAPyl in A-site of ribosome
Method: single particle / : Krahn N, Zhang J, Melnikov SV, Tharp JM, Villa A, Patel A, Howard RJ, Gabir H, Patel TR, Stetefeld J, Puglisi J, Soll D

EMDB-41709: 
Structure of C-terminal LRRK2 bound to MLi-2
Method: single particle / : Sanz-Murillo M, Villagran-Suarez A, Alegrio-Louro J, Leschziner A

EMDB-41728: 
Structure of the C-terminal half of LRRK2 bound to GZD-824 (G2019S mutant)
Method: single particle / : Villagran-Suarez A, Sanz-Murillo M, Alegrio-Louro J, Leschziner A

EMDB-41753: 
Structure of the C-terminal half of LRRK2 bound to GZD-824 (I2020T mutant)
Method: single particle / : Villagran-Suarez A, Sanz-Murillo M, Alegrio-Louro J, Leschziner A

EMDB-41754: 
Structure of C-terminal LRRK2 bound to MLi-2 (G2019S mutant)
Method: single particle / : Sanz-Murillo M, Villagran-Suarez A, Alegrio-Louro J, Leschziner A

EMDB-41756: 
Structure of C-terminal half of LRRK2 bound to GZD-824
Method: single particle / : Villagran-Suarez A, Sanz-Murillo M, Alegrio-Louro J, Leschziner A

EMDB-41757: 
Structure of full length LRRK2 bound to GZD-824 (I2020T mutant)
Method: single particle / : Villagran-Suarez A, Sanz-Murillo M, Alegrio-Louro J, Leschziner A

EMDB-41758: 
Structure of C-terminal LRRK2 bound to MLi-2 (I2020T mutant)
Method: single particle / : Sanz-Murillo M, Villagran-Suarez A, Alegrio Louro J, Leschziner A

EMDB-41759: 
Structure of full-length LRRK2 bound to MLi-2 (I2020T mutant)
Method: single particle / : Sanz-Murillo M, Villagran-Suarez A, Alegrio Louro J, Leschziner A

EMDB-41794: 
Structure of C-terminal half of LRRK2 (I2020T mutant) bound to GZD-824, Kinase-WD40
Method: single particle / : Villagran-Suarez A, Sanz-Murillo M, Alegrio-Louro J, Leschziner A

EMDB-41795: 
Structure of C-terminal half of LRRK2 (I2020T mutant) bound to GZD-824, ROC-COR domain
Method: single particle / : Villagran-Suarez A, Sanz-Murillo M, Alegrio-Louro J, Leschziner A
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
