-Search query
-Search result
Showing 1 - 50 of 379 items for (author: patel & k)

EMDB-40032: 
Structure of Trypanosoma (MDH)4-(Pex5)2, distal conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

PDB-8gh3: 
Structure of Trypanosoma (MDH)4-(Pex5)2, distal conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G
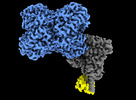
EMDB-40056: 
Structure of Trypanosoma docking complex
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

PDB-8gi0: 
Structure of Trypanosoma docking complex
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G
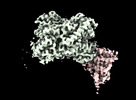
EMDB-40003: 
Structure of Trypanosoma (MDH)4-Pex5, close conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

PDB-8ggd: 
Structure of Trypanosoma (MDH)4-Pex5, close conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G
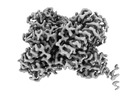
EMDB-40053: 
Consensus cryo-EM reconstruction of Trypanosoma Docking complex
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

EMDB-40008: 
Structure of Trypanosoma (MDH)4-PEX5, distal conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

PDB-8ggh: 
Structure of Trypanosoma (MDH)4-PEX5, distal conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

EMDB-40031: 
Structure of Trypanosoma (MDH)4-(Pex5)2, close conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

PDB-8gh2: 
Structure of Trypanosoma (MDH)4-(Pex5)2, close conformation
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

EMDB-40055: 
Focused map of Pex region of Trypanosoma docking complex, (MDH)4-(Pex5)1-(Pex14)1
Method: single particle / : Sonani RR, Artur B, Jemiola-Rzeminska M, Lipinski O, Patel SN, Sood T, Dubin G

EMDB-29950: 
SARS-Cov2 S protein structure in complex with neutralizing monoclonal antibody 002-S21B10
Method: single particle / : Patel A, Ortlund EA

EMDB-29975: 
Overall map of SARS-Cov2 S protein structure in complex with neutralizing monoclonal antibody 002-S21B10
Method: single particle / : Patel A, Ortlund EA

EMDB-40007: 
Local map of SARS-Cov2 S protein structure in complex with neutralizing monoclonal antibody 002-S21B10
Method: single particle / : Patel A, Ortlund EA

EMDB-42455: 
Candidatus Methanomethylophilus alvus tRNAPyl in A-site of ribosome
Method: single particle / : Krahn N, Zhang J, Melnikov SV, Tharp JM, Villa A, Patel A, Howard RJ, Gabir H, Patel TR, Stetefeld J, Puglisi J, Soll D

PDB-8upt: 
Candidatus Methanomethylophilus alvus tRNAPyl in A-site of ribosome
Method: single particle / : Krahn N, Zhang J, Melnikov SV, Tharp JM, Villa A, Patel A, Howard RJ, Gabir H, Patel TR, Stetefeld J, Puglisi J, Soll D

EMDB-40477: 
KLHDC2 in complex with EloB and EloC
Method: single particle / : Digianantonio KM, Bekes M

PDB-8sh2: 
KLHDC2 in complex with EloB and EloC
Method: single particle / : Digianantonio KM, Bekes M
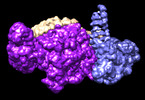
EMDB-41097: 
cryoEM structure of Smc5/6 5mer
Method: single particle / : Yu Y, Patel DJ

EMDB-41098: 
Smc5/6 8mer
Method: single particle / : Yu Y, Patel DJ

EMDB-29281: 
Cryo-EM structure of STING oligomer bound to cGAMP and NVS-STG2
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

EMDB-29282: 
Cryo-EM structure of STING oligomer bound to cGAMP, NVS-STG2 and C53
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

PDB-8flk: 
Cryo-EM structure of STING oligomer bound to cGAMP and NVS-STG2
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

PDB-8flm: 
Cryo-EM structure of STING oligomer bound to cGAMP, NVS-STG2 and C53
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

EMDB-15662: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 4-mer RNA, pppGpGpUpA (IC4)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

EMDB-15664: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 5-mer RNA, pppGpGpApApA (IC5)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

EMDB-15665: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 6-mer RNA, pppGpGpApApApU (IC6)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

EMDB-16442: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 7-mer RNA, pppGpGpUpApApApU (IC7)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

EMDB-16443: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 8-mer RNA, pppGpGpUpApApApUpG (IC8)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

EMDB-18183: 
Cryo-EM structure of IC8', a second state of yeast mitochondrial RNA polymerase transcription initiation complex with 8-mer RNA, pppGpGpUpApApApUpG
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

PDB-8att: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 4-mer RNA, pppGpGpUpA (IC4)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

PDB-8atv: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 5-mer RNA, pppGpGpApApA (IC5)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

PDB-8atw: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 6-mer RNA, pppGpGpApApApU (IC6)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

PDB-8c5s: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 7-mer RNA, pppGpGpUpApApApU (IC7)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

PDB-8c5u: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with 8-mer RNA, pppGpGpUpApApApUpG (IC8)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

PDB-8q63: 
Cryo-EM structure of IC8', a second state of yeast mitochondrial RNA polymerase transcription initiation complex with 8-mer RNA, pppGpGpUpApApApUpG
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

EMDB-15556: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with two GTP molecules poised for de novo initiation (IC2)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

PDB-8ap1: 
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex with two GTP molecules poised for de novo initiation (IC2)
Method: single particle / : Goovaerts Q, Shen J, Patel SS, Das K

EMDB-29700: 
Cryo-EM imaging scaffold subunits A and B used to display KRAS G12C complex with GDP
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

EMDB-29713: 
KRAS G12C complex with GDP imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

EMDB-29715: 
KRAS G12C complex with GDP and AMG 510 imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

EMDB-29718: 
Green Fluorescence Protein imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

EMDB-29719: 
KRAS G12V complex with GDP imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO
EMDB-29720: 
KRAS G13C complex with GDP imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

PDB-8g3k: 
Cryo-EM imaging scaffold subunits A and B used to display KRAS G12C complex with GDP
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

PDB-8g42: 
KRAS G12C complex with GDP imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO

PDB-8g47: 
KRAS G12C complex with GDP and AMG 510 imaged on a cryo-EM imaging scaffold
Method: single particle / : Castells-Graells R, Sawaya MR, Yeates TO
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
