+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8t8f | ||||||
|---|---|---|---|---|---|---|---|
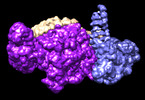
| Title | Smc5/6 8mer | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  DNA BINDING PROTEIN / Smc / Nse / DNA BINDING PROTEIN / Smc / Nse /  Condensin / Condensin /  Cohesin / Cohesin /  DNA binding / DNA binding /  DNA damage / DNA damage /  DNA repair DNA repair | ||||||
| Function / homology |  Function and homology information Function and homology informationSmc5-Smc6 complex / resolution of DNA recombination intermediates / DNA double-strand break attachment to nuclear envelope / chromosome separation / SUMOylation of DNA damage response and repair proteins / chromatin looping /  recombinational repair / recombinational repair /  regulation of telomere maintenance / protein sumoylation / double-strand break repair via homologous recombination ...Smc5-Smc6 complex / resolution of DNA recombination intermediates / DNA double-strand break attachment to nuclear envelope / chromosome separation / SUMOylation of DNA damage response and repair proteins / chromatin looping / regulation of telomere maintenance / protein sumoylation / double-strand break repair via homologous recombination ...Smc5-Smc6 complex / resolution of DNA recombination intermediates / DNA double-strand break attachment to nuclear envelope / chromosome separation / SUMOylation of DNA damage response and repair proteins / chromatin looping /  recombinational repair / recombinational repair /  regulation of telomere maintenance / protein sumoylation / double-strand break repair via homologous recombination / regulation of telomere maintenance / protein sumoylation / double-strand break repair via homologous recombination /  single-stranded DNA binding / site of double-strand break / damaged DNA binding / single-stranded DNA binding / site of double-strand break / damaged DNA binding /  chromosome, telomeric region / chromosome, telomeric region /  DNA repair / DNA repair /  ATP hydrolysis activity / ATP hydrolysis activity /  mitochondrion / mitochondrion /  ATP binding / ATP binding /  nucleus / nucleus /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Saccharomyces cerevisiae W303 (yeast) Saccharomyces cerevisiae W303 (yeast) | ||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 4.8 Å cryo EM / Resolution: 4.8 Å | ||||||
 Authors Authors | Yu, Y. / Patel, D.J. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Molecular basis for Nse5-6 mediated regulation of Smc5/6 functions. Authors: Shibai Li / You Yu / Jian Zheng / Victoria Miller-Browne / Zheng Ser / Huihui Kuang / Dinshaw J Patel / Xiaolan Zhao /   Abstract: The Smc5/6 complex (Smc5/6) is important for genome replication and repair in eukaryotes. Its cellular functions are closely linked to the ATPase activity of the Smc5 and Smc6 subunits. This activity ...The Smc5/6 complex (Smc5/6) is important for genome replication and repair in eukaryotes. Its cellular functions are closely linked to the ATPase activity of the Smc5 and Smc6 subunits. This activity requires the dimerization of the motor domains of the two SMC subunits and is regulated by the six non-SMC subunits (Nse1 to Nse6). Among the NSEs, Nse5 and Nse6 form a stable subcomplex (Nse5-6) that dampens the ATPase activity of the complex. However, the underlying mechanisms and biological significance of this regulation remain unclear. Here, we address these issues using structural and functional studies. We determined cryo-EM structures of the yeast Smc5/6 derived from complexes consisting of either all eight subunits or a subset of five subunits. Both structures reveal that Nse5-6 associates with Smc6's motor domain and the adjacent coiled-coil segment, termed the neck region. Our structural analyses reveal that this binding is compatible with motor domain dimerization but results in dislodging the Nse4 subunit from the Smc6 neck. As the Nse4-Smc6 neck interaction favors motor domain engagement and thus ATPase activity, Nse6's competition with Nse4 can explain how Nse5-6 disfavors ATPase activity. Such regulation could in principle differentially affect Smc5/6-mediated processes depending on their needs of the complex's ATPase activity. Indeed, mutagenesis data in cells provide evidence that the Nse6-Smc6 neck interaction is important for the resolution of DNA repair intermediates but not for replication termination. Our results thus provide a molecular basis for how Nse5-6 modulates the ATPase activity and cellular functions of Smc5/6. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8t8f.cif.gz 8t8f.cif.gz | 340.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8t8f.ent.gz pdb8t8f.ent.gz | 249.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8t8f.json.gz 8t8f.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/t8/8t8f https://data.pdbj.org/pub/pdb/validation_reports/t8/8t8f ftp://data.pdbj.org/pub/pdb/validation_reports/t8/8t8f ftp://data.pdbj.org/pub/pdb/validation_reports/t8/8t8f | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  41098MC  8t8eC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein |  / Killer toxin-resistance protein 29 / Killer toxin-resistance protein 29Mass: 53836.758 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Saccharomyces cerevisiae W303 (yeast) / Production host: Saccharomyces cerevisiae W303 (yeast) / Production host:   Saccharomyces cerevisiae W303 (yeast) / References: UniProt: P40026 Saccharomyces cerevisiae W303 (yeast) / References: UniProt: P40026 |
|---|---|
| #2: Protein | Mass: 64079.609 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Saccharomyces cerevisiae W303 (yeast) / Production host: Saccharomyces cerevisiae W303 (yeast) / Production host:   Saccharomyces cerevisiae W303 (yeast) / References: UniProt: Q03718 Saccharomyces cerevisiae W303 (yeast) / References: UniProt: Q03718 |
| #3: Protein | Mass: 126236.180 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Details: Smc5 E1015Q mutation, visible part is from 32-246aa and 883-1093aa. Source: (gene. exp.)   Saccharomyces cerevisiae W303 (yeast) / Production host: Saccharomyces cerevisiae W303 (yeast) / Production host:   Escherichia coli BL21(DE3) (bacteria) / References: UniProt: Q08204 Escherichia coli BL21(DE3) (bacteria) / References: UniProt: Q08204 |
| #4: Protein | Mass: 128198.742 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Details: Smc6 E1048Q mutation with C terminal 2X strep tag, visible part is from 79-286aa and 923-1084aa Source: (gene. exp.)   Saccharomyces cerevisiae W303 (yeast) / Production host: Saccharomyces cerevisiae W303 (yeast) / Production host:   Escherichia coli BL21(DE3) (bacteria) / References: UniProt: Q12749 Escherichia coli BL21(DE3) (bacteria) / References: UniProt: Q12749 |
| #5: Protein | Mass: 46195.945 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Saccharomyces cerevisiae W303 (yeast) / Production host: Saccharomyces cerevisiae W303 (yeast) / Production host:   Escherichia coli BL21(DE3) (bacteria) / References: UniProt: P43124 Escherichia coli BL21(DE3) (bacteria) / References: UniProt: P43124 |
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Smc5/6 8mer / Type: COMPLEX / Entity ID: all / Source: RECOMBINANT |
|---|---|
| Molecular weight | Value: 400 kDa/nm / Experimental value: NO |
| Source (natural) | Organism:   Saccharomyces cerevisiae W303 (yeast) Saccharomyces cerevisiae W303 (yeast) |
| Source (recombinant) | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Buffer solution | pH: 7.5 / Details: 25 mM Hepes, pH 7.5, 250 mM NaCl, 1 mM DTT |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
| Specimen support | Grid material: GOLD / Grid mesh size: 300 divisions/in. / Grid type: UltrAuFoil R1.2/1.3 |
Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2500 nm / Nominal defocus min: 1000 nm Bright-field microscopy / Nominal defocus max: 2500 nm / Nominal defocus min: 1000 nm |
| Image recording | Electron dose: 53.55 e/Å2 / Film or detector model: GATAN K3 (6k x 4k) |
- Processing
Processing
CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
|---|---|
3D reconstruction | Resolution: 4.8 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 145290 / Symmetry type: POINT |
 Movie
Movie Controller
Controller



 PDBj
PDBj