-Search query
-Search result
Showing all 48 items for (author: langlois & r)

EMDB-17705: 
Structure of Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class I
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17706: 
Structure of Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class II
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17707: 
Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class III
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17709: 
Structure of Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class V
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17710: 
Structure of Chelator-GIDSR4 - Fbp1 - phospho-Ubc8~ubiquitin - class IV
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17713: 
Catalytic module of human CTLH E3 ligase bound to multiphosphorylated UBE2H~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Prabu RJ, Schulman BA

EMDB-17715: 
SRS and Cat modules of human CTLHSR4 bound to multiphosphorylated UBE2H~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-17716: 
Structure of CTLHSR4 - phospho-UBE2H~ubiquitin bound to engineered VH
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA
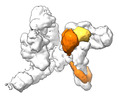
EMDB-17717: 
SRS and Cat modules of yeast Chelator-GIDSR4 bound to multiphosphorylated Ubc8~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA
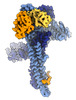
EMDB-17764: 
Catalytic module of yeast GID E3 ligase bound to multiphosphorylated Ubc8~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Prabu RJ, Schulman BA

PDB-8pjn: 
Catalytic module of human CTLH E3 ligase bound to multiphosphorylated UBE2H~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Prabu RJ, Schulman BA

PDB-8pmq: 
Catalytic module of yeast GID E3 ligase bound to multiphosphorylated Ubc8~ubiquitin
Method: single particle / : Chrustowicz J, Sherpa D, Prabu RJ, Schulman BA

EMDB-12537: 
Structure of human CTLH SR4 complex
Method: single particle / : Sherpa D, Chrustowicz J, Schulman BA

EMDB-12538: 
Structure of endogenous yeast GID Ant complex
Method: single particle / : Sherpa D, Chrustowicz J, Qiao S, Schulman BA

EMDB-12540: 
Structure of endogenous yeast Chelator-GID Ant complex
Method: single particle / : Sherpa D, Chrustowicz J, Qiao S, Schulman BA

EMDB-12541: 
Structure of Apo Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Sherpa D, Chrustowicz J, Qiao S, Prabu JR, Schulman BA

EMDB-12542: 
Structure of human CTLH-WDR26 supramolecular assembly
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-12545: 
Structure of supramolecular assembly and substrate receptor scaffolding modules of human CTLH-WDR26 complex
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-12547: 
Structure of supramolecular assembly and substrate receptor scaffolding modules of human CTLH-MKLN1 complex
Method: single particle / : Chrustowicz J, Sherpa D, Schulman BA

EMDB-12548: 
Structure of GID SR4 complex
Method: single particle / : Sherpa D, Chrustowicz J, Schulman BA

EMDB-12557: 
Structure of yeast Chelator-GID SR4 E3 ubiquitin ligase bound to its tetrameric metabolic enzyme substrate Fbp1
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

EMDB-12559: 
Substrate receptor scaffolding module of yeast Chelator-GID SR4 E3 ubiquitin ligase bound to Fbp1 substrate
Method: single particle / : Sherpa D, Chrustowicz J, Prabu JR, Schulman BA

EMDB-12560: 
Catalytic module of yeast Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Sherpa D, Chrustowicz J, Prabu JR, Schulman BA

EMDB-12563: 
Supramolecular assembly module of yeast Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

EMDB-12564: 
Substrate receptor scaffolding module of human CTLH E3 ubiquitin ligase
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

PDB-7ns3: 
Substrate receptor scaffolding module of yeast Chelator-GID SR4 E3 ubiquitin ligase bound to Fbp1 substrate
Method: single particle / : Sherpa D, Chrustowicz J, Prabu JR, Schulman BA

PDB-7ns4: 
Catalytic module of yeast Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Sherpa D, Chrustowicz J, Prabu JR, Schulman BA

PDB-7nsb: 
Supramolecular assembly module of yeast Chelator-GID SR4 E3 ubiquitin ligase
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

PDB-7nsc: 
Substrate receptor scaffolding module of human CTLH E3 ubiquitin ligase
Method: single particle / : Chrustowicz J, Sherpa D, Prabu JR, Schulman BA

EMDB-10326: 
Structure of inactive GID E3 ubiquitin ligase complex
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10327: 
Structure of active GID E3 ubiquitin ligase complex
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10328: 
Structure of GID Scaffold subcomplex
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10329: 
Structure of GID Scaffold subcomplex bound to substrate receptor Gid10
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10330: 
Structure of GID Scaffold Subcomplex bound to substrate receptor Gid4
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10331: 
Structure of endogenous inactive GID E3 ubiquitin ligase complex
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10332: 
Structure of active GID E3 ubiquitin ligase complex with RING domains deletion
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-10333: 
Structure of active GID E3 ubiquitin ligase complex minus Gid2 and delta Gid9 RING domain
Method: single particle / : Qiao S, Prabu JR, Schulman BA

PDB-6swy: 
Structure of active GID E3 ubiquitin ligase complex minus Gid2 and delta Gid9 RING domain
Method: single particle / : Qiao S, Prabu JR, Schulman BA

EMDB-3405: 
Mammalian 80S Ribosomes Associate with a Novel Vesicular Organelle
Method: subtomogram averaging / : Carter SD, Hampton CM, Langlois R, Grassucci RA, Farino ZJ, Morgenstern TM, Rice WJ, Velasco KR, Wigge C, Xu Y, Koller A, Melero R, Mitchell WG, Yi E, Aguilar JI, Levy ES, Greenberg NL, Li W, Courel M, Mahata SK, Freyberg R, Javitch JA, Di Paolo G, Chen EI, Chan RB, Carazo JM, Area-Gomez E, Jensen GJ, Frank J, Freyberg Z

EMDB-6315: 
Activation of GTP Hydrolysis in mRNA-tRNA Translocation by Elongation Factor G
Method: single particle / : Li W, Liu Z, Koripella RK, Langlois R, Sanyal S, Frank J

EMDB-6316: 
Activation of GTP Hydrolysis in mRNA-tRNA Translocation by Elongation Factor G
Method: single particle / : Li W, Liu Z, Koripella RK, Sanyal S, Frank J

PDB-3j9z: 
Activation of GTP Hydrolysis in mRNA-tRNA Translocation by Elongation Factor G
Method: single particle / : Li W, Liu Z, Koripella RK, Langlois R, Sanyal S, Frank J

PDB-3ja1: 
Activation of GTP Hydrolysis in mRNA-tRNA Translocation by Elongation Factor G
Method: single particle / : Li W, Liu Z, Koripella RK, Langlois R, Sanyal S, Frank J

EMDB-6044: 
Ribosome conformation along minimum free-energy trajectory
Method: single particle / : Dashti A, Schwander P, Langlois R, Fung R, Li W, Hosseinizadeh A, Liao HY, Pallesen J, Sharma G, Stupina VA, Simon AE, Dinman J, Frank J, Ourmazd A

EMDB-2450: 
Cryo-EM map of the CSFV IRES in complex with the small ribosomal 40S subunit and DHX29
Method: single particle / : Hashem Y, des Georges A, Dhote A, Langlois R, Liao HY, Grassucci RA, Pestova TV, Hellen CUT, Frank J

EMDB-2451: 
Cryo-EM structure of the CSFV IRES in complex with eIF3, small ribosomal 40S subunit and DHX29
Method: single particle / : Hashem Y, des-Georges A, Dhote V, Langlois R, Liao HY, Grassucci RA, Pestova TV, Hellen CUT, Frank J

PDB-4c4q: 
Cryo-EM map of the CSFV IRES in complex with the small ribosomal 40S subunit and DHX29
Method: single particle / : Hashem Y, desGeorges A, Dhote V, Langlois R, Liao HY, Grassucci RA, Pestova TV, Hellen CUT, Frank J

EMDB-5658: 
Cryo-electron microscopy structure of the mammalian 43S preinitiation complex bound to DHX29
Method: single particle / : Hashem Y, des Georges A, Dhote V, Langlois R, Liao HY, Grassucci RA, Hellen CUT, Pestova TV, Frank J
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
