[English] 日本語
 Yorodumi
Yorodumi- EMDB-17764: Catalytic module of yeast GID E3 ligase bound to multiphosphoryla... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 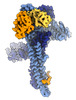 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Catalytic module of yeast GID E3 ligase bound to multiphosphorylated Ubc8~ubiquitin | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | E3 ubiquitin ligase / E2 ubiquitin-conjugating enzyme / phosphorylation / GID / LIGASE | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationGID complex / Regulation of pyruvate metabolism / E2 ubiquitin-conjugating enzyme / ubiquitin conjugating enzyme activity / negative regulation of gluconeogenesis / Maturation of protein E / Maturation of protein E / ER Quality Control Compartment (ERQC) / Myoclonic epilepsy of Lafora / FLT3 signaling by CBL mutants ...GID complex / Regulation of pyruvate metabolism / E2 ubiquitin-conjugating enzyme / ubiquitin conjugating enzyme activity / negative regulation of gluconeogenesis / Maturation of protein E / Maturation of protein E / ER Quality Control Compartment (ERQC) / Myoclonic epilepsy of Lafora / FLT3 signaling by CBL mutants / Prevention of phagosomal-lysosomal fusion / IRAK2 mediated activation of TAK1 complex / Alpha-protein kinase 1 signaling pathway / Glycogen synthesis / Regulation of TBK1, IKKε (IKBKE)-mediated activation of IRF3, IRF7 / IRAK1 recruits IKK complex / IRAK1 recruits IKK complex upon TLR7/8 or 9 stimulation / Membrane binding and targetting of GAG proteins / Regulation of TBK1, IKKε-mediated activation of IRF3, IRF7 upon TLR3 ligation / Endosomal Sorting Complex Required For Transport (ESCRT) / IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation / PTK6 Regulates RTKs and Their Effectors AKT1 and DOK1 / Negative regulation of FLT3 / Constitutive Signaling by NOTCH1 HD Domain Mutants / Regulation of FZD by ubiquitination / TICAM1,TRAF6-dependent induction of TAK1 complex / TICAM1-dependent activation of IRF3/IRF7 / NOTCH2 Activation and Transmission of Signal to the Nucleus / APC/C:Cdc20 mediated degradation of Cyclin B / p75NTR recruits signalling complexes / Downregulation of ERBB4 signaling / NOTCH3 Activation and Transmission of Signal to the Nucleus / APC-Cdc20 mediated degradation of Nek2A / TRAF6-mediated induction of TAK1 complex within TLR4 complex / TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling / PINK1-PRKN Mediated Mitophagy / Pexophagy / Regulation of innate immune responses to cytosolic DNA / InlA-mediated entry of Listeria monocytogenes into host cells / VLDLR internalisation and degradation / Regulation of pyruvate metabolism / Downregulation of ERBB2:ERBB3 signaling / NF-kB is activated and signals survival / Regulation of PTEN localization / NRIF signals cell death from the nucleus / Regulation of BACH1 activity / Activated NOTCH1 Transmits Signal to the Nucleus / Synthesis of active ubiquitin: roles of E1 and E2 enzymes / Translesion synthesis by REV1 / MAP3K8 (TPL2)-dependent MAPK1/3 activation / Translesion synthesis by POLK / TICAM1, RIP1-mediated IKK complex recruitment / Downregulation of TGF-beta receptor signaling / EGFR downregulation / Activation of IRF3, IRF7 mediated by TBK1, IKKε (IKBKE) / Translesion synthesis by POLI / Gap-filling DNA repair synthesis and ligation in GG-NER / Josephin domain DUBs / Regulation of activated PAK-2p34 by proteasome mediated degradation / InlB-mediated entry of Listeria monocytogenes into host cell / IKK complex recruitment mediated by RIP1 / JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 / TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) / N-glycan trimming in the ER and Calnexin/Calreticulin cycle / TNFR1-induced NF-kappa-B signaling pathway / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / Asymmetric localization of PCP proteins / TCF dependent signaling in response to WNT / SCF-beta-TrCP mediated degradation of Emi1 / NIK-->noncanonical NF-kB signaling / Ubiquitin-dependent degradation of Cyclin D / Regulation of NF-kappa B signaling / AUF1 (hnRNP D0) binds and destabilizes mRNA / TNFR2 non-canonical NF-kB pathway / activated TAK1 mediates p38 MAPK activation / Vpu mediated degradation of CD4 / Assembly of the pre-replicative complex / Negative regulators of DDX58/IFIH1 signaling / Deactivation of the beta-catenin transactivating complex / Degradation of DVL / CDK-mediated phosphorylation and removal of Cdc6 / Ubiquitin Mediated Degradation of Phosphorylated Cdc25A / Regulation of signaling by CBL / Dectin-1 mediated noncanonical NF-kB signaling / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / Fanconi Anemia Pathway / Hh mutants are degraded by ERAD / Negative regulation of FGFR3 signaling / SCF(Skp2)-mediated degradation of p27/p21 / Degradation of AXIN / Peroxisomal protein import / Recognition of DNA damage by PCNA-containing replication complex / Degradation of GLI1 by the proteasome / Activation of NF-kappaB in B cells / Regulation of TNFR1 signaling / Negative regulation of FGFR2 signaling / Downregulation of SMAD2/3:SMAD4 transcriptional activity / Stabilization of p53 / Termination of translesion DNA synthesis Similarity search - Function | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.53 Å | ||||||||||||
 Authors Authors | Chrustowicz J / Sherpa D / Prabu RJ / Schulman BA | ||||||||||||
| Funding support |  Germany, European Union, 3 items Germany, European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2024 Journal: Mol Cell / Year: 2024Title: Multisite phosphorylation dictates selective E2-E3 pairing as revealed by Ubc8/UBE2H-GID/CTLH assemblies. Authors: Jakub Chrustowicz / Dawafuti Sherpa / Jerry Li / Christine R Langlois / Eleftheria C Papadopoulou / D Tung Vu / Laura A Hehl / Özge Karayel / Viola Beier / Susanne von Gronau / Judith ...Authors: Jakub Chrustowicz / Dawafuti Sherpa / Jerry Li / Christine R Langlois / Eleftheria C Papadopoulou / D Tung Vu / Laura A Hehl / Özge Karayel / Viola Beier / Susanne von Gronau / Judith Müller / J Rajan Prabu / Matthias Mann / Gary Kleiger / Arno F Alpi / Brenda A Schulman /   Abstract: Ubiquitylation is catalyzed by coordinated actions of E3 and E2 enzymes. Molecular principles governing many important E3-E2 partnerships remain unknown, including those for RING-family GID/CTLH E3 ...Ubiquitylation is catalyzed by coordinated actions of E3 and E2 enzymes. Molecular principles governing many important E3-E2 partnerships remain unknown, including those for RING-family GID/CTLH E3 ubiquitin ligases and their dedicated E2, Ubc8/UBE2H (yeast/human nomenclature). GID/CTLH-Ubc8/UBE2H-mediated ubiquitylation regulates biological processes ranging from yeast metabolic signaling to human development. Here, cryoelectron microscopy (cryo-EM), biochemistry, and cell biology reveal this exquisitely specific E3-E2 pairing through an unconventional catalytic assembly and auxiliary interactions 70-100 Å away, mediated by E2 multisite phosphorylation. Rather than dynamic polyelectrostatic interactions reported for other ubiquitylation complexes, multiple Ubc8/UBE2H phosphorylation sites within acidic CK2-targeted sequences specifically anchor the E2 C termini to E3 basic patches. Positions of phospho-dependent interactions relative to the catalytic domains correlate across evolution. Overall, our data show that phosphorylation-dependent multivalency establishes a specific E3-E2 partnership, is antagonistic with dephosphorylation, rigidifies the catalytic centers within a flexing GID E3-substrate assembly, and facilitates substrate collision with ubiquitylation active sites. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17764.map.gz emd_17764.map.gz | 5.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17764-v30.xml emd-17764-v30.xml emd-17764.xml emd-17764.xml | 23.4 KB 23.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17764_fsc.xml emd_17764_fsc.xml | 9.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_17764.png emd_17764.png | 108 KB | ||
| Masks |  emd_17764_msk_1.map emd_17764_msk_1.map | 76.8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17764.cif.gz emd-17764.cif.gz | 6.9 KB | ||
| Others |  emd_17764_additional_1.map.gz emd_17764_additional_1.map.gz emd_17764_half_map_1.map.gz emd_17764_half_map_1.map.gz emd_17764_half_map_2.map.gz emd_17764_half_map_2.map.gz | 66.8 MB 60 MB 60 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17764 http://ftp.pdbj.org/pub/emdb/structures/EMD-17764 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17764 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17764 | HTTPS FTP |
-Validation report
| Summary document |  emd_17764_validation.pdf.gz emd_17764_validation.pdf.gz | 685.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_17764_full_validation.pdf.gz emd_17764_full_validation.pdf.gz | 685.4 KB | Display | |
| Data in XML |  emd_17764_validation.xml.gz emd_17764_validation.xml.gz | 16.7 KB | Display | |
| Data in CIF |  emd_17764_validation.cif.gz emd_17764_validation.cif.gz | 22.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17764 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17764 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17764 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-17764 | HTTPS FTP |
-Related structure data
| Related structure data |  8pmqMC  8pjnC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17764.map.gz / Format: CCP4 / Size: 76.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17764.map.gz / Format: CCP4 / Size: 76.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.427 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17764_msk_1.map emd_17764_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Map post-processed with DeepEMhancer
| File | emd_17764_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map post-processed with DeepEMhancer | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_17764_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17764_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of yeast Chelator-GIDSR4 E3 ligase, tetrameric Fbp1 subst...
| Entire | Name: Complex of yeast Chelator-GIDSR4 E3 ligase, tetrameric Fbp1 substrate and multiphosphorylated Ubc8~ubiquitin |
|---|---|
| Components |
|
-Supramolecule #1: Complex of yeast Chelator-GIDSR4 E3 ligase, tetrameric Fbp1 subst...
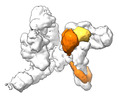
| Supramolecule | Name: Complex of yeast Chelator-GIDSR4 E3 ligase, tetrameric Fbp1 substrate and multiphosphorylated Ubc8~ubiquitin type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 Details: Map obtained by focus refinement over the catalytic module (Gid2, Gid9) and Ubc8~ubiquitin |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.7 MDa |
-Macromolecule #1: E3 ubiquitin-protein ligase RMD5
| Macromolecule | Name: E3 ubiquitin-protein ligase RMD5 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: RING-type E3 ubiquitin transferase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 49.244594 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSELLDSFET EFAKFYTDSN LEETNLQKCL DHTHEFKSQL KKLKAHLNKH IQESKPEVYN KLSDKEKQKF KRKRELIIEK LSKSQRQWD HSVKKQIKYV SQQSNRFNKS TLNKLKEFDI DSVYVNKLPK ETMENVNEAI GYHILRYSID NMPLGNKNEA F QYLKDVYG ...String: MSELLDSFET EFAKFYTDSN LEETNLQKCL DHTHEFKSQL KKLKAHLNKH IQESKPEVYN KLSDKEKQKF KRKRELIIEK LSKSQRQWD HSVKKQIKYV SQQSNRFNKS TLNKLKEFDI DSVYVNKLPK ETMENVNEAI GYHILRYSID NMPLGNKNEA F QYLKDVYG ITNKESTEFI EMGQIVHDLK KGDTESCLKW CSNEMESLSS NHTALSSLKF DLYTLSAMQI VKHGNPVELY YQ ITQNAPL DCFRHREKEL MQNVVPLLTK SLIGQPIEDI DSKVNKELKE CTSLFIKEYC AAKHIFFDSP LFLIVLSGLI SFQ FFIKYK TIRELAHVDW TTKDELPFDV KLPDFLTHFH PIFICPVLKE ETTTENPPYS LACHHIISKK ALDRLSKNGT ITFK CPYCP VNTSMSSTKK VRFVML UniProtKB: E3 ubiquitin-protein ligase RMD5 |
-Macromolecule #2: Protein FYV10
| Macromolecule | Name: Protein FYV10 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: RING-type E3 ubiquitin transferase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 59.975102 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MAEKSIFNEP DVDFHLKLNQ QLFHIPYELL SKRIKHTQAV INKETKSLHE HTAALNQIFE HNDVEHDELA LAKITEMIRK VDHIERFLN TQIKSYCQIL NRIKKRLEFF HELKDIKSQN SGTSHNGNNE GTRTKLIQWY QSYTNILIGD YLTRNNPIKY N SETKDHWN ...String: MAEKSIFNEP DVDFHLKLNQ QLFHIPYELL SKRIKHTQAV INKETKSLHE HTAALNQIFE HNDVEHDELA LAKITEMIRK VDHIERFLN TQIKSYCQIL NRIKKRLEFF HELKDIKSQN SGTSHNGNNE GTRTKLIQWY QSYTNILIGD YLTRNNPIKY N SETKDHWN SGVVFLKQSQ LDDLIDYDVL LEANRISTSL LHERNLLPLI SWINENKKTL TKKSSILEFQ ARLQEYIELL KV DNYTDAI VCFQRFLLPF VKSNFTDLKL ASGLLIFIKY CNDQKPTSST SSGFDTEEIK SQSLPMKKDR IFQHFFHKSL PRI TSKPAV NTTDYDKSSL INLQSGDFER YLNLLDDQRW SVLNDLFLSD FYSMYGISQN DPLLIYLSLG ISSLKTRDCL HPSD DENGN QETETATTAE KEVEDLQLFT LHSLKRKNCP VCSETFKPIT QALPFAHHIQ SQLFENPILL PNGNVYDSKK LKKLA KTLK KQNLISLNPG QIMDPVDMKI FCESDSIKMY PT UniProtKB: Protein FYV10 |
-Macromolecule #3: Ubiquitin-conjugating enzyme E2-24 kDa
| Macromolecule | Name: Ubiquitin-conjugating enzyme E2-24 kDa / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: E2 ubiquitin-conjugating enzyme |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.920771 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSSSKRRIET DVMKLLMSDH QVDLINDSMQ EFHVKFLGPK DTPYENGVWR LHVELPDNYP YKSPSIGFVN KIFHPNIDIA SGSIKLDVI NSTWSPLYDL INIVEWMIPG LLKEPNGSDP LNNEAATLQL RDKKLYEEKI KEYIDKYATK EKYQQMFGGD N DSDDSDSG ...String: MSSSKRRIET DVMKLLMSDH QVDLINDSMQ EFHVKFLGPK DTPYENGVWR LHVELPDNYP YKSPSIGFVN KIFHPNIDIA SGSIKLDVI NSTWSPLYDL INIVEWMIPG LLKEPNGSDP LNNEAATLQL RDKKLYEEKI KEYIDKYATK EKYQQMFGGD N DSDDSDSG GDLQEEDSDS DEDMDGTGVS SGDDSVDEL(SEP) EDL(SEP)DIDV(SEP)D DDDYDEVANQ UniProtKB: Ubiquitin-conjugating enzyme E2-24 kDa |
-Macromolecule #4: Ubiquitin
| Macromolecule | Name: Ubiquitin / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.576831 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQIFVKTLTG KTITLEVEPS DTIENVKAKI QDKEGIPPDQ QRLIFAGKQL EDGRTLSDYN IQKESTLHLV LRLRGG UniProtKB: Polyubiquitin-C |
-Macromolecule #5: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 69.24 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




























 Z
Z Y
Y X
X



































