-Search query
-Search result
Showing 1 - 50 of 460 items for (author: deng & b)
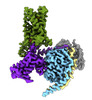
EMDB-43647: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL
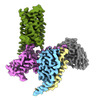
EMDB-43650: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-43656: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-43657: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)
Method: single particle / : Kim Y, Gumpper RH, Roth BL

PDB-8vy7: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

PDB-8vy9: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-40650: 
Phosphoinositide phosphate 3 kinase gamma
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40651: 
Phosphoinositide phosphate 3 kinase gamma bound with ATP
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40652: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40653: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and Gbetagamma
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40654: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and two Gbetagamma subunits in State 1
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-40655: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and two Gbetagamma subunits in State 2
Method: single particle / : Chen CL, Tesmer JJG, Bandekar SJ, Cash J

EMDB-43091: 
hGBP1 conformer on the bacterial outer membrane
Method: subtomogram averaging / : Zhu S, MacMicking J

EMDB-43153: 
hGBP1 conformer on the bacterial outer membrane
Method: subtomogram averaging / : Zhu S, MacMicking J

EMDB-35832: 
Cryo-EM structure of the GPR34 receptor in complex with the antagonist YL-365
Method: single particle / : Jia GW, Wang X, Zhang CB, Dong HH, Su ZM

PDB-8iyx: 
Cryo-EM structure of the GPR34 receptor in complex with the antagonist YL-365
Method: single particle / : Jia GW, Wang X, Zhang CB, Dong HH, Su ZM

EMDB-37516: 
BA.2(S375) Spike (S6P)/hACE2 complex
Method: single particle / : Wei X, Zhang Z

EMDB-37517: 
Local refinement of RBD-ACE2
Method: single particle / : Wei X, Zhang Z

EMDB-43542: 
ELIC5 with cysteamine in 2:1:1 POPC:POPE:POPG nanodisc in open conformation
Method: single particle / : Petroff II JT, Deng Z, Rau MJ, Fitzpatrick JAJ, Yuan P, Cheng WWL

PDB-8vuw: 
ELIC5 with cysteamine in 2:1:1 POPC:POPE:POPG nanodisc in open conformation
Method: single particle / : Petroff II JT, Deng Z, Rau MJ, Fitzpatrick JAJ, Yuan P, Cheng WWL

EMDB-36368: 
Cryo-EM structure of CCHFV envelope protein Gc trimer in complex with Gc13 Fab
Method: single particle / : Chong T, Cao S

EMDB-36406: 
CCHFV envelope protein Gc in complex with Gc8
Method: single particle / : Chong T, Cao S

EMDB-36407: 
CCHFV envelope protein Gc in complex with Gc13
Method: single particle / : Chong T, Cao S

PDB-8jkd: 
Cryo-EM structure of CCHFV envelope protein Gc trimer in complex with Gc13 Fab
Method: single particle / : Chong T, Cao S

EMDB-34552: 
Structure of dimeric mouse SCMC core complex
Method: single particle / : Chi P, Ou G, Han Z, Li J, Deng D
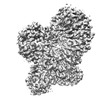
EMDB-34554: 
Structure of mouse SCMC bound with KH domain of FILIA
Method: single particle / : Chi P, Ou G, Han Z, Li J, Deng D

EMDB-34555: 
Structure of mouse SCMC bound with full-length FILIA
Method: single particle / : Chi P, Ou G, Han Z, Li L, Deng D

EMDB-34556: 
Structure of mouse SCMC core complex
Method: single particle / : Chi P, Ou G, Li J, Han Z, Deng D

PDB-8h93: 
Structure of dimeric mouse SCMC core complex
Method: single particle / : Chi P, Ou G, Han Z, Li J, Deng D

PDB-8h94: 
Structure of mouse SCMC bound with KH domain of FILIA
Method: single particle / : Chi P, Ou G, Han Z, Li J, Deng D

PDB-8h95: 
Structure of mouse SCMC bound with full-length FILIA
Method: single particle / : Chi P, Ou G, Han Z, Li L, Deng D

PDB-8h96: 
Structure of mouse SCMC core complex
Method: single particle / : Chi P, Ou G, Li J, Han Z, Deng D

EMDB-40270: 
Cryo-EM structure of GPR34-Gi complex
Method: single particle / : Yong XH, Zhao C, Yan W, Shao ZH

PDB-8sai: 
Cryo-EM structure of GPR34-Gi complex
Method: single particle / : Yong XH, Zhao C, Yan W, Shao ZH

EMDB-34954: 
Cryo-EM structure of SSX1 bound to the H2AK119Ub nucleosome at a resolution of 3.05 angstrom
Method: single particle / : Zebin T, Ai HS, Ziyu X, GuoChao C, Man P, Liu L

EMDB-36747: 
Cryo-EM structure of 2 SSX1 bound to H2AK119Ub nucleosome at a resolution of 3.6 angstroms (2:1 complex)
Method: single particle / : Zebin T, Ai HS, Ziyu X, GuoChao C, Man P, Liu L

PDB-8hqy: 
Cryo-EM structure of SSX1 bound to the H2AK119Ub nucleosome at a resolution of 3.05 angstrom
Method: single particle / : Zebin T, Ai HS, Ziyu X, GuoChao C, Man P, Liu L

EMDB-36751: 
Outward_facing SLC15A4 monomer
Method: single particle / : Zhang SS, Chen XD, Xie M

EMDB-36752: 
Outward-facing SLC15A4 dimer
Method: single particle / : Zhang SS, Chen XD, Xie M

EMDB-36753: 
SLC15A4_TASL complex
Method: single particle / : Zhang SS, Chen XD, Xie M

EMDB-35347: 
Structure of R2 with 3'UTR and DNA in binding state
Method: single particle / : Deng P, Tan S, Wang J, Liu JJ

EMDB-35348: 
Structure of R2 with 3'UTR and DNA in unwinding state
Method: single particle / : Deng P, Tan S, Wang J, Liu JJ

EMDB-35349: 
Structure of R2 with 5'ORF
Method: single particle / : Deng P, Tan S, Wang J, Liu JJ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search










 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
