[English] 日本語
 Yorodumi
Yorodumi- PDB-8hqy: Cryo-EM structure of SSX1 bound to the H2AK119Ub nucleosome at a ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8hqy | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
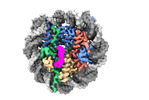
| Title | Cryo-EM structure of SSX1 bound to the H2AK119Ub nucleosome at a resolution of 3.05 angstrom | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords |  STRUCTURAL PROTEIN / SSX1 / H2AK119Ub nucleosome / Synovial Sarcoma / ssBAF / STRUCTURAL PROTEIN / SSX1 / H2AK119Ub nucleosome / Synovial Sarcoma / ssBAF /  reader protein reader protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationprotein localization to CENP-A containing chromatin / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere ...protein localization to CENP-A containing chromatin / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere / Meiotic synapsis / RNA Polymerase I Promoter Opening / Assembly of the ORC complex at the origin of replication /  DNA methylation / Condensation of Prophase Chromosomes / HCMV Late Events / Chromatin modifications during the maternal to zygotic transition (MZT) / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / SIRT1 negatively regulates rRNA expression / DNA methylation / Condensation of Prophase Chromosomes / HCMV Late Events / Chromatin modifications during the maternal to zygotic transition (MZT) / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / SIRT1 negatively regulates rRNA expression /  innate immune response in mucosa / PRC2 methylates histones and DNA / Defective pyroptosis / HDACs deacetylate histones / RNA Polymerase I Promoter Escape / Nonhomologous End-Joining (NHEJ) / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / NoRC negatively regulates rRNA expression / G2/M DNA damage checkpoint / B-WICH complex positively regulates rRNA expression / DNA Damage/Telomere Stress Induced Senescence / Metalloprotease DUBs / RMTs methylate histone arginines / innate immune response in mucosa / PRC2 methylates histones and DNA / Defective pyroptosis / HDACs deacetylate histones / RNA Polymerase I Promoter Escape / Nonhomologous End-Joining (NHEJ) / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / NoRC negatively regulates rRNA expression / G2/M DNA damage checkpoint / B-WICH complex positively regulates rRNA expression / DNA Damage/Telomere Stress Induced Senescence / Metalloprotease DUBs / RMTs methylate histone arginines /  Meiotic recombination / Pre-NOTCH Transcription and Translation / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / UCH proteinases / Meiotic recombination / Pre-NOTCH Transcription and Translation / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / UCH proteinases /  nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / E3 ubiquitin ligases ubiquitinate target proteins / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / RUNX1 regulates transcription of genes involved in differentiation of HSCs / chromatin organization / Processing of DNA double-strand break ends / HATs acetylate histones / antibacterial humoral response / Senescence-Associated Secretory Phenotype (SASP) / Oxidative Stress Induced Senescence / killing of cells of another organism / Estrogen-dependent gene expression / defense response to Gram-negative bacterium / Ub-specific processing proteases / defense response to Gram-positive bacterium / Amyloid fiber formation / protein heterodimerization activity / negative regulation of cell population proliferation / regulation of DNA-templated transcription / nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / E3 ubiquitin ligases ubiquitinate target proteins / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / RUNX1 regulates transcription of genes involved in differentiation of HSCs / chromatin organization / Processing of DNA double-strand break ends / HATs acetylate histones / antibacterial humoral response / Senescence-Associated Secretory Phenotype (SASP) / Oxidative Stress Induced Senescence / killing of cells of another organism / Estrogen-dependent gene expression / defense response to Gram-negative bacterium / Ub-specific processing proteases / defense response to Gram-positive bacterium / Amyloid fiber formation / protein heterodimerization activity / negative regulation of cell population proliferation / regulation of DNA-templated transcription /  DNA binding / DNA binding /  extracellular space / extracellular exosome / extracellular space / extracellular exosome /  nucleoplasm / nucleoplasm /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.05 Å cryo EM / Resolution: 3.05 Å | |||||||||
 Authors Authors | Zebin, T. / Ai, H.S. / Ziyu, X. / GuoChao, C. / Man, P. / Liu, L. | |||||||||
| Funding support |  China, 2items China, 2items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Synovial sarcoma X breakpoint 1 protein uses a cryptic groove to selectively recognize H2AK119Ub nucleosomes. Authors: Zebin Tong / Huasong Ai / Ziyu Xu / Kezhang He / Guo-Chao Chu / Qiang Shi / Zhiheng Deng / Qiaomei Xue / Maoshen Sun / Yunxiang Du / Lujun Liang / Jia-Bin Li / Man Pan / Lei Liu /  Abstract: The cancer-specific fusion oncoprotein SS18-SSX1 disturbs chromatin accessibility by hijacking the BAF complex from the promoters and enhancers to the Polycomb-repressed chromatin regions. This ...The cancer-specific fusion oncoprotein SS18-SSX1 disturbs chromatin accessibility by hijacking the BAF complex from the promoters and enhancers to the Polycomb-repressed chromatin regions. This process relies on the selective recognition of H2AK119Ub nucleosomes by synovial sarcoma X breakpoint 1 (SSX1). However, the mechanism underlying the selective recognition of H2AK119Ub nucleosomes by SSX1 in the absence of ubiquitin (Ub)-binding capacity remains unknown. Here we report the cryo-EM structure of SSX1 bound to H2AK119Ub nucleosomes at 3.1-Å resolution. Combined in vitro biochemical and cellular assays revealed that the Ub recognition by SSX1 is unique and depends on a cryptic basic groove formed by H3 and the Ub motif on the H2AK119 site. Moreover, this unorthodox binding mode of SSX1 induces DNA unwrapping at the entry/exit sites. Together, our results describe a unique mode of site-specific ubiquitinated nucleosome recognition that underlies the specific hijacking of the BAF complex to Polycomb regions by SS18-SSX1 in synovial sarcoma. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8hqy.cif.gz 8hqy.cif.gz | 328 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8hqy.ent.gz pdb8hqy.ent.gz | 246.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8hqy.json.gz 8hqy.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/hq/8hqy https://data.pdbj.org/pub/pdb/validation_reports/hq/8hqy ftp://data.pdbj.org/pub/pdb/validation_reports/hq/8hqy ftp://data.pdbj.org/pub/pdb/validation_reports/hq/8hqy | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  34954MC  8hr1C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Protein , 5 types, 7 molecules AEBDHFU
| #1: Protein |  Mass: 11401.268 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: CALMAC_LOCUS17614 / Production host: Homo sapiens (human) / Gene: CALMAC_LOCUS17614 / Production host:   Escherichia coli (E. coli) / References: UniProt: A0A653DHJ5 Escherichia coli (E. coli) / References: UniProt: A0A653DHJ5#2: Protein | | Mass: 9509.197 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: N305_08114 / Production host: Homo sapiens (human) / Gene: N305_08114 / Production host:   Escherichia coli (E. coli) / References: UniProt: A0A093PN76 Escherichia coli (E. coli) / References: UniProt: A0A093PN76#4: Protein | Mass: 10076.528 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: H2BC12, H2BFT, HIRIP1, HIST1H2BK / Production host: Homo sapiens (human) / Gene: H2BC12, H2BFT, HIRIP1, HIST1H2BK / Production host:   Escherichia coli (E. coli) / References: UniProt: O60814 Escherichia coli (E. coli) / References: UniProt: O60814#5: Protein | |  Mass: 9222.823 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Escherichia coli (E. coli) / References: UniProt: A0A8C2GGI7 Escherichia coli (E. coli) / References: UniProt: A0A8C2GGI7#8: Protein | | Mass: 8462.727 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: UBB / Production host: Homo sapiens (human) / Gene: UBB / Production host:   Escherichia coli (E. coli) / References: UniProt: J3QS39 Escherichia coli (E. coli) / References: UniProt: J3QS39 |
|---|
-Histone H2A type 1- ... , 2 types, 2 molecules CG
| #3: Protein | Mass: 11740.742 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: H2AC4, H2AFM, HIST1H2AB, H2AC8, H2AFA, HIST1H2AE / Production host: Homo sapiens (human) / Gene: H2AC4, H2AFM, HIST1H2AB, H2AC8, H2AFA, HIST1H2AE / Production host:   Escherichia coli (E. coli) / References: UniProt: P04908 Escherichia coli (E. coli) / References: UniProt: P04908 |
|---|---|
| #6: Protein | Mass: 11637.600 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: HIST1H2AB, H2AFM, HIST1H2AE, H2AFA / Production host: Homo sapiens (human) / Gene: HIST1H2AB, H2AFM, HIST1H2AE, H2AFA / Production host:   Escherichia coli (E. coli) / References: UniProt: P04908 Escherichia coli (E. coli) / References: UniProt: P04908 |
-Protein/peptide , 1 types, 1 molecules S
| #7: Protein/peptide | Mass: 2912.261 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: SSX2, SSX2A, SSX2B / Production host: Homo sapiens (human) / Gene: SSX2, SSX2A, SSX2B / Production host:   Escherichia coli (E. coli) / References: UniProt: Q16385 Escherichia coli (E. coli) / References: UniProt: Q16385 |
|---|
-DNA chain , 2 types, 2 molecules IJ
| #9: DNA chain | Mass: 41800.641 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
|---|---|
| #10: DNA chain | Mass: 42151.840 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: SSX1-H2AK119Ub nucleosome complex / Type: COMPLEX / Entity ID: #1-#4, #7-#10 / Source: RECOMBINANT |
|---|---|
| Molecular weight | Value: 0.29 MDa / Experimental value: NO |
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Source (recombinant) | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Buffer solution | pH: 7.5 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2500 nm / Nominal defocus min: 1000 nm Bright-field microscopy / Nominal defocus max: 2500 nm / Nominal defocus min: 1000 nm |
| Image recording | Average exposure time: 2.56 sec. / Electron dose: 50 e/Å2 / Film or detector model: GATAN K3 (6k x 4k) |
- Processing
Processing
CTF correction | Type: NONE | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
3D reconstruction | Resolution: 3.05 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 195091 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller




 PDBj
PDBj










































