[English] 日本語
 Yorodumi
Yorodumi- EMDB-2024: Three-dimensional reconstruction of another targeted individual 1... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2024 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
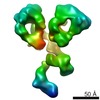
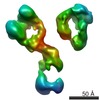
| Title | Three-dimensional reconstruction of another targeted individual 17nm nascent high-density lipoprotein (HDL) particle by individual-particle electron tomography (IPET). The reconstruction displayed a ring-shaped structure of containing three protein - apoipoprotein A-Is (28 kDa each). | |||||||||
 Map data Map data | 17nm high density lipoprotein (HDL) particle | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | reconstituted 17nm nascent high-density lipoprotein (HDL) | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | subtomogram averaging / cryo EM / negative staining / Resolution: 36.0 Å | |||||||||
 Authors Authors | Zhang L / Ren G | |||||||||
 Citation Citation |  Journal: PLoS One / Year: 2012 Journal: PLoS One / Year: 2012Title: IPET and FETR: experimental approach for studying molecular structure dynamics by cryo-electron tomography of a single-molecule structure. Authors: Lei Zhang / Gang Ren /  Abstract: The dynamic personalities and structural heterogeneity of proteins are essential for proper functioning. Structural determination of dynamic/heterogeneous proteins is limited by conventional ...The dynamic personalities and structural heterogeneity of proteins are essential for proper functioning. Structural determination of dynamic/heterogeneous proteins is limited by conventional approaches of X-ray and electron microscopy (EM) of single-particle reconstruction that require an average from thousands to millions different molecules. Cryo-electron tomography (cryoET) is an approach to determine three-dimensional (3D) reconstruction of a single and unique biological object such as bacteria and cells, by imaging the object from a series of tilting angles. However, cconventional reconstruction methods use large-size whole-micrographs that are limited by reconstruction resolution (lower than 20 Å), especially for small and low-symmetric molecule (<400 kDa). In this study, we demonstrated the adverse effects from image distortion and the measuring tilt-errors (including tilt-axis and tilt-angle errors) both play a major role in limiting the reconstruction resolution. Therefore, we developed a "focused electron tomography reconstruction" (FETR) algorithm to improve the resolution by decreasing the reconstructing image size so that it contains only a single-instance protein. FETR can tolerate certain levels of image-distortion and measuring tilt-errors, and can also precisely determine the translational parameters via an iterative refinement process that contains a series of automatically generated dynamic filters and masks. To describe this method, a set of simulated cryoET images was employed; to validate this approach, the real experimental images from negative-staining and cryoET were used. Since this approach can obtain the structure of a single-instance molecule/particle, we named it individual-particle electron tomography (IPET) as a new robust strategy/approach that does not require a pre-given initial model, class averaging of multiple molecules or an extended ordered lattice, but can tolerate small tilt-errors for high-resolution single "snapshot" molecule structure determination. Thus, FETR/IPET provides a completely new opportunity for a single-molecule structure determination, and could be used to study the dynamic character and equilibrium fluctuation of macromolecules. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2024.map.gz emd_2024.map.gz | 28.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2024-v30.xml emd-2024-v30.xml emd-2024.xml emd-2024.xml | 10.7 KB 10.7 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2024.tif EMD-2024.tif | 48.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2024 http://ftp.pdbj.org/pub/emdb/structures/EMD-2024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2024 | HTTPS FTP |
-Validation report
| Summary document |  emd_2024_validation.pdf.gz emd_2024_validation.pdf.gz | 210.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2024_full_validation.pdf.gz emd_2024_full_validation.pdf.gz | 209.8 KB | Display | |
| Data in XML |  emd_2024_validation.xml.gz emd_2024_validation.xml.gz | 6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2024 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2024 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2024 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2024 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2024.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2024.map.gz / Format: CCP4 / Size: 29.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 17nm high density lipoprotein (HDL) particle | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.73 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : second 17nm nascent HDL particle
| Entire | Name: second 17nm nascent HDL particle |
|---|---|
| Components |
|
-Supramolecule #1000: second 17nm nascent HDL particle
| Supramolecule | Name: second 17nm nascent HDL particle / type: sample / ID: 1000 Details: Fresh sample was prepared with 2 days before using. Oligomeric state: monomer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 200 KDa / Theoretical: 200 KDa |
-Macromolecule #1: 17 nm nascent HDL
| Macromolecule | Name: 17 nm nascent HDL / type: protein_or_peptide / ID: 1 / Name.synonym: 17 nm nascent HDL Details: HDL contains lipids and three apoA-I molecules. Each A-I molecular weight is about 28kDa. Number of copies: 1 / Oligomeric state: monomer / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Strain: nascent HDL / synonym: Human / Location in cell: Plasma Homo sapiens (human) / Strain: nascent HDL / synonym: Human / Location in cell: Plasma |
| Molecular weight | Experimental: 200 KDa / Theoretical: 200 KDa |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
- Sample preparation
Sample preparation
| Concentration | 0.01 mg/mL |
|---|---|
| Buffer | pH: 7.4 Details: 1X Dulbeccos phosphate-buffered saline (Invitrogen, La Jolla, CA), 2.7 mM KCl, 1.46 mM KH2PO4, 136.9 mM NaCl, and 8.1 mM Na2HPO4 |
| Staining | Type: NEGATIVE Details: EM Specimens were prepared by standard cryo-EM protocol. |
| Grid | Details: 200 mesh glow-discharged holey thin carbon-coated EM grids (Cu-200HN, Pacific Grid-Tech, USA) |
| Vitrification | Cryogen name: NITROGEN / Chamber humidity: 90 % / Chamber temperature: 103 K / Instrument: HOMEMADE PLUNGER / Details: Vitrification instrument: Manual / Method: Blot for 2 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Temperature | Average: 100 K |
| Details | Tilt step is 1.5 degree |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Sampling interval: 1.73 µm / Number real images: 93 / Average electron dose: 140 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.2 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 67000 |
| Sample stage | Specimen holder: Gatan / Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Min angle: -69 ° / Tilt series - Axis1 - Max angle: 69 ° |
- Image processing
Image processing
| Details | Single targeted particle"s images were reconstructed by focus ET reconstruction (FETR) algorithm of individual particle electron tomography (IPET) method. Average number of tilts used in the 3D reconstructions: 93. Average tomographic tilt angle increment: 1.5. |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 36.0 Å / Resolution method: OTHER / Software - Name: IPET, and, FETR Details: Map was reconstructed by individual-particle electron tomography (IPET)and Focus ET Reconstruction Algorithm. |
| CTF correction | Details: TOMOCTF |
| Final angle assignment | Details: Tomography tilt angle from -60 to 60 in step of 1.5 |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera