+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3kaz | ||||||
|---|---|---|---|---|---|---|---|
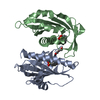
| Title | Crystal structure of abscisic acid receptor PYL2 | ||||||
 Components Components | Putative uncharacterized protein At2g26040 | ||||||
 Keywords Keywords |  SIGNALING PROTEIN / phytohormone receptor / PYR/PYL/RCAR / abscisic acid signaling SIGNALING PROTEIN / phytohormone receptor / PYR/PYL/RCAR / abscisic acid signaling | ||||||
| Function / homology |  Function and homology information Function and homology informationprotein phosphatase inhibitor complex /  abscisic acid binding / abscisic acid-activated signaling pathway / protein phosphatase inhibitor activity / abscisic acid binding / abscisic acid-activated signaling pathway / protein phosphatase inhibitor activity /  signaling receptor activity / protein homodimerization activity / identical protein binding / signaling receptor activity / protein homodimerization activity / identical protein binding /  nucleus / nucleus /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.85 Å molecular replacement / Resolution: 1.85 Å | ||||||
 Authors Authors | Zhou, X.E. / Melcher, K. / Ng, L.-M. / Soon, F.-F. / Xu, Y. / Suino-Powell, K.M. / Kovach, A. / Li, J. / Xu, H.E. | ||||||
 Citation Citation |  Journal: Nature / Year: 2009 Journal: Nature / Year: 2009Title: Agate-latch-lock mechanism for hormone signalling by abscisic acid receptors Authors: Melcher, K. / Ng, L.-M. / Zhou, X.E. / Soon, F.-F. / Xu, Y. / Suino-Powell, K.-M. / Park, S.-Y. / Weiner, J.J. / Fujii, H. / Chinnusamy, V. / Kovach, A. / Li, J. / Wang, Y. / Li, J.Y. / ...Authors: Melcher, K. / Ng, L.-M. / Zhou, X.E. / Soon, F.-F. / Xu, Y. / Suino-Powell, K.-M. / Park, S.-Y. / Weiner, J.J. / Fujii, H. / Chinnusamy, V. / Kovach, A. / Li, J. / Wang, Y. / Li, J.Y. / Peterson, F.C. / Jensen, D.R. / Yong, E.-L. / Volkman, B.F. / Cutler, S.R. / Zhu, J.-K. / Xu, H.E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3kaz.cif.gz 3kaz.cif.gz | 224.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3kaz.ent.gz pdb3kaz.ent.gz | 183.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3kaz.json.gz 3kaz.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ka/3kaz https://data.pdbj.org/pub/pdb/validation_reports/ka/3kaz ftp://data.pdbj.org/pub/pdb/validation_reports/ka/3kaz ftp://data.pdbj.org/pub/pdb/validation_reports/ka/3kaz | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 19848.361 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Arabidopsis thaliana (thale cress) / Gene: At2g26040 / Plasmid: pET24a / Production host: Arabidopsis thaliana (thale cress) / Gene: At2g26040 / Plasmid: pET24a / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: O80992 Escherichia coli (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: O80992#2: Chemical |  1,3-Butanediol 1,3-Butanediol#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.54 Å3/Da / Density % sol: 51.56 % |
|---|---|
Crystal grow | Temperature: 298 K / Method: vapor diffusion / pH: 8.2 Details: Potassium phosphate dibasic, pH 8.2, VAPOR DIFFUSION, temperature 298K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 21-ID-D / Wavelength: 1 Å / Beamline: 21-ID-D / Wavelength: 1 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Aug 16, 2009 |
| Radiation | Monochromator: Ni FILTER / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1 Å / Relative weight: 1 : 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.85→30 Å / Num. all: 52118 / Num. obs: 49719 / % possible obs: 92.6 % / Observed criterion σ(F): 2 / Observed criterion σ(I): 2 / Redundancy: 11.4 % / Rsym value: 0.085 |
-Phasing
Phasing | Method:  molecular replacement molecular replacement | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Phasing MR | Model details: Phaser MODE: MR_AUTO
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT / Resolution: 1.85→29.48 Å / Cor.coef. Fo:Fc: 0.952 / Cor.coef. Fo:Fc free: 0.938 / Occupancy max: 1 / Occupancy min: 1 / SU B: 7.528 / SU ML: 0.103 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE MOLECULAR REPLACEMENT / Resolution: 1.85→29.48 Å / Cor.coef. Fo:Fc: 0.952 / Cor.coef. Fo:Fc free: 0.938 / Occupancy max: 1 / Occupancy min: 1 / SU B: 7.528 / SU ML: 0.103 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 48.15 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.85→29.48 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.85→1.9 Å
|
 Movie
Movie Controller
Controller















 PDBj
PDBj


