+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1tc3 | ||||||
|---|---|---|---|---|---|---|---|
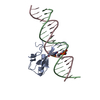
| Title | TRANSPOSASE TC3A1-65 FROM CAENORHABDITIS ELEGANS | ||||||
 Components Components |
| ||||||
 Keywords Keywords | DNA BINDING PROTEIN/DNA / TRANSPOSASE / DNA BINDING / HELIX-TURN-HELIX / TC1/MARINER FAMILY / COMPLEX (TRANSPOSASE-DNA) / DNA BINDING PROTEIN-DNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MIRAS / Resolution: 2.45 Å MIRAS / Resolution: 2.45 Å | ||||||
 Authors Authors | Van Pouderoyen, G. / Ketting, R.F. / Perrakis, A. / Plasterk, R.H.A. / Sixma, T.K. | ||||||
 Citation Citation |  Journal: EMBO J. / Year: 1997 Journal: EMBO J. / Year: 1997Title: Crystal structure of the specific DNA-binding domain of Tc3 transposase of C.elegans in complex with transposon DNA. Authors: van Pouderoyen, G. / Ketting, R.F. / Perrakis, A. / Plasterk, R.H. / Sixma, T.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1tc3.cif.gz 1tc3.cif.gz | 47.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1tc3.ent.gz pdb1tc3.ent.gz | 30.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1tc3.json.gz 1tc3.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1tc3_validation.pdf.gz 1tc3_validation.pdf.gz | 440.8 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1tc3_full_validation.pdf.gz 1tc3_full_validation.pdf.gz | 452.2 KB | Display | |
| Data in XML |  1tc3_validation.xml.gz 1tc3_validation.xml.gz | 8 KB | Display | |
| Data in CIF |  1tc3_validation.cif.gz 1tc3_validation.cif.gz | 10 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/tc/1tc3 https://data.pdbj.org/pub/pdb/validation_reports/tc/1tc3 ftp://data.pdbj.org/pub/pdb/validation_reports/tc/1tc3 ftp://data.pdbj.org/pub/pdb/validation_reports/tc/1tc3 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 6543.232 Da / Num. of mol.: 1 / Source method: obtained synthetically |
|---|---|
| #2: DNA chain | Mass: 6029.904 Da / Num. of mol.: 1 / Source method: obtained synthetically |
| #3: Protein | Mass: 5813.720 Da / Num. of mol.: 1 / Fragment: SPECIFIC DNA BINDING DOMAIN, RESIDUES 2 - 52 / Mutation: C-TERMINAL 6-HIS TAG Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.7 Å3/Da / Density % sol: 55 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Method: vapor diffusion, hanging drop / pH: 5.5 Details: PROTEIN/DNA COMPLEX WAS CRYSTALLIZED FROM 15% MPD, 100 MM NACL, 20 MM CACL2, 10 MM DTT, 50 MM NA ACETATE PH 5.5, VAPOR DIFFUSION, HANGING DROP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal | *PLUS Density % sol: 55 % | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 4 ℃ | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 113 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  EMBL/DESY, HAMBURG EMBL/DESY, HAMBURG  / Beamline: BW7A / Beamline: BW7A |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Aug 1, 1996 / Details: BW7A SYSTEM |
| Radiation | Monochromator: BW7A SYSTEM / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Relative weight: 1 |
| Reflection | Resolution: 2.45→30 Å / Num. obs: 8089 / % possible obs: 89.7 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 8.8 % / Biso Wilson estimate: 58.5 Å2 / Rmerge(I) obs: 0.055 / Rsym value: 0.055 / Net I/σ(I): 19.4 |
| Reflection shell | Resolution: 2.45→2.51 Å / Redundancy: 3 % / Rmerge(I) obs: 0.504 / Mean I/σ(I) obs: 1.93 / Rsym value: 0.504 / % possible all: 67 |
| Reflection | *PLUS Highest resolution: 2.45 Å / Lowest resolution: 30 Å / % possible obs: 89.7 % / Num. measured all: 70943 |
| Reflection shell | *PLUS % possible obs: 67.1 % / Rmerge(I) obs: 0.438 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIRAS / Resolution: 2.45→20 Å / Isotropic thermal model: TNT BCORREL / Cross valid method: R FREE THROUGHOUT / σ(F): 0 / Stereochemistry target values: TNT PROTGEO MIRAS / Resolution: 2.45→20 Å / Isotropic thermal model: TNT BCORREL / Cross valid method: R FREE THROUGHOUT / σ(F): 0 / Stereochemistry target values: TNT PROTGEODetails: X-PLOR AND REFMAC/ARP WERE USED IN EARLIER STAGES OF REFINEMENT.
| ||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: TRONRUD ET AL. / Bsol: 150 Å2 / ksol: 0.75 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.45→20 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: TNT / Version: 5-E / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.45 Å / Lowest resolution: 20 Å / σ(F): 0 / % reflection Rfree: 5 % / Rfactor Rwork: 0.234 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller











 PDBj
PDBj







































