+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 9dp9 | ||||||
|---|---|---|---|---|---|---|---|
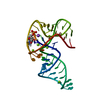
| Title | Structure of a Tick-Borne Flavivirus xrRNA | ||||||
 Components Components | RNA (440-MER) | ||||||
 Keywords Keywords | RNA / Virus / xrRNA / Exonuclease | ||||||
| Function / homology | RNA / RNA (> 10) / RNA (> 100) Function and homology information Function and homology information | ||||||
| Biological species |  Powassan virus Powassan virus | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 4.1 Å | ||||||
 Authors Authors | Langeberg, C.J. / Kieft, J.S. / Sherlock, M.E. / Szucs, M.J. / Vicens, Q. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Nat Commun / Year: 2025 Journal: Nat Commun / Year: 2025Title: Tick-borne flavivirus exoribonuclease-resistant RNAs contain a double loop structure. Authors: Conner J Langeberg / Matthew J Szucs / Madeline E Sherlock / Quentin Vicens / Jeffrey S Kieft /  Abstract: Viruses from the Flaviviridae family contain human relevant pathogens that generate subgenomic noncoding RNAs during infection using structured exoribonuclease resistant RNAs (xrRNAs). These xrRNAs ...Viruses from the Flaviviridae family contain human relevant pathogens that generate subgenomic noncoding RNAs during infection using structured exoribonuclease resistant RNAs (xrRNAs). These xrRNAs block progression of host cell's 5' to 3' exoribonucleases. The structures of several xrRNAs from mosquito-borne and insect-specific flaviviruses reveal a conserved fold in which a ring-like motif encircles the 5' end of the xrRNA. However, the xrRNAs found in tick-borne and no known vector flaviviruses have distinct characteristics, and their 3-D fold was unsolved. Here, we verify the presence of xrRNAs in the encephalitis-causing tick-borne Powassan Virus. We characterize their secondary structure and obtain a mid-resolution map of one of these xrRNAs using cryo-EM, revealing a unique double-loop ring element. Integrating these results with covariation analysis, biochemical data, and existing high-resolution structural information yields a model in which the core of the fold matches the previously solved xrRNA fold, but the expanded double loop ring is remodeled upon encountering the exoribonuclease. These results are representative of a broad class of xrRNAs and reveal a conserved strategy of structure-based exoribonuclease resistance achieved through a unique topology across a viral family of importance to global health. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  9dp9.cif.gz 9dp9.cif.gz | 222.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb9dp9.ent.gz pdb9dp9.ent.gz | 165.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  9dp9.json.gz 9dp9.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dp/9dp9 https://data.pdbj.org/pub/pdb/validation_reports/dp/9dp9 ftp://data.pdbj.org/pub/pdb/validation_reports/dp/9dp9 ftp://data.pdbj.org/pub/pdb/validation_reports/dp/9dp9 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  47099MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: RNA chain | Mass: 142224.047 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Powassan virus / Variant: Spooner isolate Powassan virus / Variant: Spooner isolateProduction host: in vitro transcription vector pT7-TP(deltai) (others) | ||||
|---|---|---|---|---|---|
| #2: Chemical | ChemComp-MG / Has ligand of interest | N | Has protein modification | N | |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Structure of a Tick-Borne Flavivirus xrRNA / Type: COMPLEX / Entity ID: #1 / Source: RECOMBINANT |
|---|---|
| Molecular weight | Value: 0.14244 MDa / Experimental value: NO |
| Source (natural) | Organism:  Powassan virus Powassan virus |
| Source (recombinant) | Organism: in vitro transcription vector pT7-TP(deltai) (others) |
| Buffer solution | pH: 7.5 |
| Specimen | Conc.: 3 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Specimen support | Grid material: COPPER / Grid mesh size: 400 divisions/in. / Grid type: C-flat-1.2/1.3 |
| Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 4 K |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 105000 X / Nominal defocus max: 2000 nm / Nominal defocus min: 800 nm / Cs: 2.7 mm / Alignment procedure: COMA FREE |
| Specimen holder | Cryogen: NITROGEN / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Image recording | Electron dose: 50 e/Å2 / Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) |
- Processing
Processing
| EM software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Type: PHASE FLIPPING ONLY | ||||||||||||||||||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 1061023 | ||||||||||||||||||||||||||||||||||||||||
| Symmetry | Point symmetry: C1 (asymmetric) | ||||||||||||||||||||||||||||||||||||||||
| 3D reconstruction | Resolution: 4.1 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 55485 / Num. of class averages: 1 / Symmetry type: POINT | ||||||||||||||||||||||||||||||||||||||||
| Atomic model building | B value: 150.22 / Protocol: FLEXIBLE FIT / Space: REAL | ||||||||||||||||||||||||||||||||||||||||
| Atomic model building |
|
 Movie
Movie Controller
Controller



 PDBj
PDBj