+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8f2y | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of an alternating AT dodecamer: 5'-CGCGATATCGCG-3 | ||||||||||||||||||||||||||||
 Components Components | DNA (5'-D(* Keywords KeywordsDNA / AT sequence / Minor groove | Function / homology | DNA / DNA (> 10) |  Function and homology information Function and homology informationBiological species |  Homo sapiens (human) Homo sapiens (human)Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.78 Å MOLECULAR REPLACEMENT / Resolution: 1.78 Å  Authors AuthorsOgbonna, E.N. / Wilson, W.D. | Funding support | |  United States, 1items United States, 1items
 Citation Citation Journal: Acs Bio Med Chem Au / Year: 2023 Journal: Acs Bio Med Chem Au / Year: 2023Title: X-ray Structure Characterization of the Selective Recognition of AT Base Pair Sequences. Authors: Ogbonna, E.N. / Paul, A. / Farahat, A.A. / Terrell, J.R. / Mineva, E. / Ogbonna, V. / Boykin, D.W. / Wilson, W.D. #1:  Journal: Biochemistry / Year: 1989 Journal: Biochemistry / Year: 1989Title: Molecular structure of the netropsin-d(CGCGATATCGCG) complex: DNA conformation in an alternating AT segment. Authors: Coll, M. / Aymami, J. / van der Marel, G.A. / van Boom, J.H. / Rich, A. / Wang, A.H. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8f2y.cif.gz 8f2y.cif.gz | 38.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8f2y.ent.gz pdb8f2y.ent.gz | 21.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8f2y.json.gz 8f2y.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/f2/8f2y https://data.pdbj.org/pub/pdb/validation_reports/f2/8f2y ftp://data.pdbj.org/pub/pdb/validation_reports/f2/8f2y ftp://data.pdbj.org/pub/pdb/validation_reports/f2/8f2y | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  8ec1C  8ed6C  8edaC  8edbC  8f1sC  8f1vC  8f20C  8f2wC  8f94C  8fb4C  8fdpC  8fdqC  8fdrC  1bnaS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 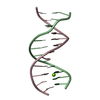
| |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||||||||||||||||||||
| Unit cell |
| |||||||||||||||||||||||||||
| Noncrystallographic symmetry (NCS) | NCS domain:
NCS domain segments:
NCS oper: (Code: givenMatrix: (0.710580337659, -0.702920432949, -0.0312801642587), (-0.703505296372, -0.710554893073, -0.0138579185359), (-0.0124852596732, 0.0318529256598, -0.999414583353)Vector: 19. ...NCS oper: (Code: given Matrix: (0.710580337659, -0.702920432949, -0.0312801642587), Vector: |
- Components
Components
| #1: DNA chain | Mass: 3663.392 Da / Num. of mol.: 2 / Source method: obtained synthetically Details: Native alternating AT DNA dodecamer (5'-CGCGATATCGCG-3'). Source: (synth.)  Homo sapiens (human) Homo sapiens (human)#2: Chemical | ChemComp-MG / | #3: Water | ChemComp-HOH / | Has ligand of interest | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.34 Å3/Da / Density % sol: 47.46 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 7 Details: MPD, SODIUM CACODYLATE TRIHYDRATE, SPERMINE TETRACHLORIDE, SODIUM CHLROIDE, MAGNESIUM HEXAHYDRATE |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS-II NSLS-II  / Beamline: 17-ID-1 / Wavelength: 1 Å / Beamline: 17-ID-1 / Wavelength: 1 Å |
| Detector | Type: DECTRIS EIGER X 9M / Detector: PIXEL / Date: Sep 28, 2022 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.78→25.71 Å / Num. obs: 7017 / % possible obs: 99.6 % / Redundancy: 2 % / Biso Wilson estimate: 21.95 Å2 / CC1/2: 0.994 / CC star: 0.999 / Net I/σ(I): 11.2 |
| Reflection shell | Resolution: 1.78→1.84 Å / Rmerge(I) obs: 0.1557 / Num. unique obs: 692 / CC1/2: 0.955 / Rpim(I) all: 0.1557 / Rrim(I) all: 0.2202 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1BNA Resolution: 1.78→25.71 Å / SU ML: 0.2132 / Cross valid method: FREE R-VALUE / σ(F): 1.36 / Phase error: 26.3698 Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2
| ||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 23.55 Å2 | ||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.78→25.71 Å
| ||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | Type: Torsion NCS / Rms dev position: 2.05791582552 Å | ||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller



 PDBj
PDBj









































