+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8dgz | ||||||
|---|---|---|---|---|---|---|---|
| Title | Caspase-7 bound to substrate mimic and allosteric inhibitor | ||||||
 Components Components |
| ||||||
 Keywords Keywords | APOPTOSIS/APOPTOSIS INHIBITOR / allostery / inhibitor / ternary / APOPTOSIS-APOPTOSIS INHIBITOR complex | ||||||
| Function / homology |  Function and homology information Function and homology informationlymphocyte apoptotic process / caspase-7 / positive regulation of plasma membrane repair / cellular response to staurosporine / SMAC, XIAP-regulated apoptotic response / plasma membrane repair / Activation of caspases through apoptosome-mediated cleavage / SMAC (DIABLO) binds to IAPs / SMAC(DIABLO)-mediated dissociation of IAP:caspase complexes / fibroblast apoptotic process ...lymphocyte apoptotic process / caspase-7 / positive regulation of plasma membrane repair / cellular response to staurosporine / SMAC, XIAP-regulated apoptotic response / plasma membrane repair / Activation of caspases through apoptosome-mediated cleavage / SMAC (DIABLO) binds to IAPs / SMAC(DIABLO)-mediated dissociation of IAP:caspase complexes / fibroblast apoptotic process / ceramide biosynthetic process / execution phase of apoptosis / Apoptotic cleavage of cellular proteins / Caspase-mediated cleavage of cytoskeletal proteins / response to UV / striated muscle cell differentiation / cysteine-type peptidase activity / protein catabolic process / protein maturation / protein processing / fibrillar center / peptidase activity / positive regulation of neuron apoptotic process / heart development / cellular response to lipopolysaccharide / neuron apoptotic process / aspartic-type endopeptidase activity / defense response to bacterium / cysteine-type endopeptidase activity / apoptotic process / proteolysis / : / RNA binding / nucleoplasm / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human)synthetic construct (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.8 Å MOLECULAR REPLACEMENT / Resolution: 2.8 Å | ||||||
 Authors Authors | Propp, J. / Kalenkiewicz, A. / Kathryn, F.H. / Spies, M.A. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Chemistry / Year: 2023 Journal: Chemistry / Year: 2023Title: Allosteric Tuning of Caspase-7: Establishing the Nexus of Structure and Catalytic Power. Authors: Hobbs, K.F. / Propp, J. / Vance, N.R. / Kalenkiewicz, A. / Witkin, K.R. / Ashley Spies, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8dgz.cif.gz 8dgz.cif.gz | 101.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8dgz.ent.gz pdb8dgz.ent.gz | 72.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8dgz.json.gz 8dgz.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dg/8dgz https://data.pdbj.org/pub/pdb/validation_reports/dg/8dgz ftp://data.pdbj.org/pub/pdb/validation_reports/dg/8dgz ftp://data.pdbj.org/pub/pdb/validation_reports/dg/8dgz | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  8dj3C 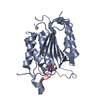 1f1jS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Unit cell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Noncrystallographic symmetry (NCS) | NCS domain:
NCS domain segments: Ens-ID: 1
|
 Movie
Movie Controller
Controller



 PDBj
PDBj


