[English] 日本語
 Yorodumi
Yorodumi- PDB-8dch: Crystal Structure of a highly resistant HIV-1 protease Clinical i... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8dch | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
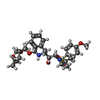
| Title | Crystal Structure of a highly resistant HIV-1 protease Clinical isolate PR10x with GRL-0519 (tris-tetrahydrofuran as P2 ligand) | |||||||||||||||
 Components Components | Protease | |||||||||||||||
 Keywords Keywords | VIRAL PROTEIN / HYDROLASE/INHIBITOR / HIV/AIDS / HIV protease / drug resistant clinical mutant / evolution of drug resistance / distal mutations / HYDROLASE-INHIBITOR complex | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationHIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding ...HIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding / viral penetration into host nucleus / host multivesicular body / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / host cell / viral nucleocapsid / DNA recombination / DNA-directed DNA polymerase / aspartic-type endopeptidase activity / Hydrolases; Acting on ester bonds / DNA-directed DNA polymerase activity / symbiont-mediated suppression of host gene expression / viral translational frameshifting / symbiont entry into host cell / lipid binding / host cell plasma membrane / host cell nucleus / virion membrane / structural molecule activity / proteolysis / DNA binding / zinc ion binding Similarity search - Function | |||||||||||||||
| Biological species |   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.25 Å molecular replacement / Resolution: 1.25 Å | |||||||||||||||
 Authors Authors | Wong-Sam, A.E. / Wang, Y.-F. / Weber, I.T. | |||||||||||||||
| Funding support |  United States, 4items United States, 4items
| |||||||||||||||
 Citation Citation |  Journal: J.Mol.Graph.Model. / Year: 2022 Journal: J.Mol.Graph.Model. / Year: 2022Title: HIV-1 protease with 10 lopinavir and darunavir resistance mutations exhibits altered inhibition, structural rearrangements and extreme dynamics. Authors: Wong-Sam, A. / Wang, Y.F. / Kneller, D.W. / Kovalevsky, A.Y. / Ghosh, A.K. / Harrison, R.W. / Weber, I.T. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8dch.cif.gz 8dch.cif.gz | 125.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8dch.ent.gz pdb8dch.ent.gz | 96.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8dch.json.gz 8dch.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dc/8dch https://data.pdbj.org/pub/pdb/validation_reports/dc/8dch ftp://data.pdbj.org/pub/pdb/validation_reports/dc/8dch ftp://data.pdbj.org/pub/pdb/validation_reports/dc/8dch | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  8dciC  3nu3S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 10784.560 Da / Num. of mol.: 2 Mutation: Q7K, L10F, I13V, L19I, K20T, V32I, L33I, E35N, M36I, M46I, G48Q, I54L, L63P, C67E, A71V, T74S, L76V, I84V, L89V, L90M, T91S, I93L, C95A Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Human immunodeficiency virus 1 / Strain: Clinical isolate PR76Vmx / Gene: gag-pol / Production host: Human immunodeficiency virus 1 / Strain: Clinical isolate PR76Vmx / Gene: gag-pol / Production host:  #2: Chemical | ChemComp-G52 / ( | #3: Chemical | #4: Chemical | ChemComp-GOL / | #5: Water | ChemComp-HOH / | Has ligand of interest | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.5 Å3/Da / Density % sol: 57.7 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 5.2 Details: Hanging-drop vapor diffusion using equal volumes of protein stock (3.9 mg/mL) and well reservoir solution. Cryoprotected in 30% glycerol. Complex on ice at 1:8 ratio of PR to PI, ...Details: Hanging-drop vapor diffusion using equal volumes of protein stock (3.9 mg/mL) and well reservoir solution. Cryoprotected in 30% glycerol. Complex on ice at 1:8 ratio of PR to PI, crystallized in 1 M NaCl and 0.1 M sodium acetate pH 5.2 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-ID / Wavelength: 1 Å / Beamline: 22-ID / Wavelength: 1 Å |
| Detector | Type: RAYONIX MX300HE / Detector: CCD / Date: Oct 25, 2017 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.25→50 Å / Num. obs: 67583 / % possible obs: 99.4 % / Redundancy: 6.8 % / Rmerge(I) obs: 0.08 / Rpim(I) all: 0.033 / Rrim(I) all: 0.086 / Χ2: 0.999 / Net I/σ(I): 20.7 / Num. measured all: 458139 |
| Reflection shell | Resolution: 1.25→1.29 Å / Redundancy: 3.3 % / Rmerge(I) obs: 0.389 / Mean I/σ(I) obs: 3.1 / Num. unique obs: 6718 / CC1/2: 0.991 / Rpim(I) all: 0.03 / Rrim(I) all: 0.078 / Χ2: 1.003 / % possible all: 97 |
-Phasing
| Phasing | Method:  molecular replacement molecular replacement | ||||||
|---|---|---|---|---|---|---|---|
| Phasing MR | R rigid body: 0.659
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3NU3 Resolution: 1.25→50 Å / Cross valid method: FREE R-VALUE / σ(F): 0 / Stereochemistry target values: ENGH AND HUBER Details: ANISOTROPIC REFINEMENT REDUCED FREE R (NO CUTOFF) BY ?
| |||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: MOEWS & KRETSINGER, J.MOL.BIOL.91(1973)201-228 | |||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 68.63 Å2 / Biso mean: 20.3406 Å2 / Biso min: 6.68 Å2 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.25→50 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller







 PDBj
PDBj