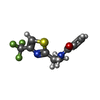| 登録情報 | データベース: PDB / ID: 7wym
|
|---|
| タイトル | Structure of the SARS-COV-2 main protease with 337 inhibitor |
|---|
 要素 要素 | 3C-like proteinase nsp5 |
|---|
 キーワード キーワード | VIRAL PROTEIN / Structure of the SARS-COV-2 main protease with 337 inhibitor |
|---|
| 機能・相同性 |  機能・相同性情報 機能・相同性情報
viral genome replication / methyltransferase activity / Assembly of the SARS-CoV-2 Replication-Transcription Complex (RTC) / Maturation of replicase proteins / ISG15-specific peptidase activity / Transcription of SARS-CoV-2 sgRNAs / Translation of Replicase and Assembly of the Replication Transcription Complex / Replication of the SARS-CoV-2 genome / double membrane vesicle viral factory outer membrane / SARS coronavirus main proteinase ...viral genome replication / methyltransferase activity / Assembly of the SARS-CoV-2 Replication-Transcription Complex (RTC) / Maturation of replicase proteins / ISG15-specific peptidase activity / Transcription of SARS-CoV-2 sgRNAs / Translation of Replicase and Assembly of the Replication Transcription Complex / Replication of the SARS-CoV-2 genome / double membrane vesicle viral factory outer membrane / SARS coronavirus main proteinase / endonuclease activity / host cell endosome / symbiont-mediated degradation of host mRNA / mRNA guanylyltransferase / symbiont-mediated suppression of host ISG15-protein conjugation / G-quadruplex RNA binding / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of IRF3 activity / methylation / omega peptidase activity / SARS-CoV-2 modulates host translation machinery / host cell Golgi apparatus / symbiont-mediated perturbation of host ubiquitin-like protein modification / ubiquitinyl hydrolase 1 / cysteine-type deubiquitinase activity / 加水分解酵素; プロテアーゼ; ペプチド結合加水分解酵素; システインプロテアーゼ / single-stranded RNA binding / regulation of autophagy / viral protein processing / host cell perinuclear region of cytoplasm / host cell endoplasmic reticulum membrane / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / symbiont-mediated suppression of host gene expression / viral translational frameshifting / symbiont-mediated activation of host autophagy / cysteine-type endopeptidase activity / lipid binding / host cell nucleus / SARS-CoV-2 activates/modulates innate and adaptive immune responses / proteolysis / zinc ion binding / membrane類似検索 - 分子機能 main proteinase (3clpro) structure, domain 3 / main proteinase (3clpro) structure, domain 3 / Trypsin-like serine proteases / Non-structural protein NSP3, SUD-N (Mac2) domain, betacoronavirus / Sarbecovirus Nsp3c-N domain profile. / Non-structural protein NSP3, N-terminal, betacoronavirus / Polyprotein cleavage domain PL2pro superfamily, betacoronavirus / Non-structural protein NSP3, SUD-N (Mac2) domain superfamily, betacoronavirus / Betacoronavirus SUD-C domain / Betacoronavirus replicase NSP3, N-terminal ...main proteinase (3clpro) structure, domain 3 / main proteinase (3clpro) structure, domain 3 / Trypsin-like serine proteases / Non-structural protein NSP3, SUD-N (Mac2) domain, betacoronavirus / Sarbecovirus Nsp3c-N domain profile. / Non-structural protein NSP3, N-terminal, betacoronavirus / Polyprotein cleavage domain PL2pro superfamily, betacoronavirus / Non-structural protein NSP3, SUD-N (Mac2) domain superfamily, betacoronavirus / Betacoronavirus SUD-C domain / Betacoronavirus replicase NSP3, N-terminal / NSP1 globular domain superfamily, betacoronavirus / Non-structural protein 2, SARS-CoV-like / Betacoronavirus single-stranded poly(A) binding domain / Betacoronavirus Nsp3c-M domain profile. / NSP1, globular domain, betacoronavirus / Non-structural protein NSP3, SUD-M domain, betacoronavirus / Non-structural protein NSP3, SUD-M domain superfamily, betacoronavirus / NSP1, C-terminal domain, betacoronavirus / Betacoronavirus replicase NSP1 / Betacoronavirus (BetaCoV) Nsp1 C-terminal domain profile. / : / Betacoronavirus Nsp3e group 2-specific marker (G2M) domain profile. / Betacoronavirus Nsp3c-C domain profile. / Betacoronavirus Nsp3e nucleic acid-binding (NAB) domain profile. / DPUP/SUD, C-terminal, betacoronavirus / Non-structural protein NSP3, nucleic acid-binding domain, betacoronavirus / Non-structural protein NSP3A domain-like superfamily / Non-structural protein NSP3, nucleic acid-binding domain superfamily, betacoronavirus / Non-structural protein 6, betacoronavirus / Betacoronavirus nucleic acid-binding (NAB) / Papain-like viral protease, palm and finger domains, coronavirus / Papain-like protease, N-terminal domain superfamily, coronavirus / Carbamoyl-phosphate synthase subdomain signature 2. / Coronavirus replicase NSP2, N-terminal / : / Coronavirus replicase NSP2, C-terminal / Coronavirus (CoV) Nsp2 middle domain profile. / NSP1, globular domain, alpha/betacoronavirus / Coronavirus (CoV) Nsp1 globular domain profile. / Coronavirus (CoV) Nsp2 N-terminal domain profile. / Coronavirus (CoV) Nsp2 C-terminal domain profile. / Nonstructural protein 2, N-terminal domain, coronavirus / Non-structural protein 2, C-terminal domain, coronavirus / NSP3, second ubiquitin-like (Ubl) domain, coronavirus / Coronavirus Nsp3a Ubl domain profile. / Coronavirus Nsp3d Ubl domain profile. / NSP3, first ubiquitin-like (Ubl) domain, coronavirus / Peptidase family C16 domain profile. / : / : / Coronavirus replicase NSP7 / Coronavirus 3Ecto domain profile. / Coronavirus RNA-dependent RNA polymerase (RdRp) Nsp7 cofactor domain profile. / Coronavirus RNA-dependent RNA polymerase (RdRp) Nsp8 cofactor domain profile. / Coronavirus Nsp9 single-stranded RNA (ssRNA)-binding domain profile. / Coronavirus (CoV) ExoN/MTase coactivator domain profile. / Coronavirus (CoV) Nsp3 Y domain profile. / Coronavirus Nsp4 C-terminal (Nsp4C) domain profile. / Coronavirus main protease (M-pro) domain profile. / Peptidase C30, coronavirus / Peptidase C16, coronavirus / Non-structural protein NSP9, coronavirus / Non-structural protein NSP7, coronavirus / Non-structural protein NSP8, coronavirus / RNA synthesis protein NSP10, coronavirus / Non-structural protein NSP4, C-terminal, coronavirus / RNA synthesis protein NSP10 superfamily, coronavirus / Non-structural protein NSP9 superfamily, coronavirus / Non-structural protein NSP7 superfamily, coronavirus / Non-structural protein NSP8 superfamily, coronavirus / Non-structural protein NSP4, C-terminal superfamily, coronavirus / Papain-like protease, thumb domain superfamily, coronavirus / Peptidase C30, domain 3, coronavirus / Non-structural protein 6, coronavirus / Coronavirus replicase NSP3, C-terminal / Non-structural protein NSP4, N-terminal, coronavirus / Coronavirus endopeptidase C30 / Coronavirus papain-like peptidase / Coronavirus replicase NSP8 / Coronavirus RNA synthesis protein NSP10 / Coronavirus replicase NSP4, C-terminal / Coronavirus replicase NSP6 / Coronavirus replicase NSP4, N-terminal / Coronavirus replicase NSP3, C-terminal / Coronavirus replicase NSP9 / Thrombin, subunit H / Non-structural protein 3, X-domain-like / Appr-1"-p processing enzyme / Macro domain / Macro domain profile. / Macro domain / Macro domain-like / Peptidase S1, PA clan, chymotrypsin-like fold / Peptidase S1, PA clan / Beta Barrel / Orthogonal Bundle / Mainly Beta / Mainly Alpha類似検索 - ドメイン・相同性 |
|---|
| 生物種 |   Severe acute respiratory syndrome coronavirus 2 (ウイルス) Severe acute respiratory syndrome coronavirus 2 (ウイルス) |
|---|
| 手法 |  X線回折 / X線回折 /  シンクロトロン / シンクロトロン /  分子置換 / 解像度: 2.05 Å 分子置換 / 解像度: 2.05 Å |
|---|
 データ登録者 データ登録者 | Qin, B. / Hou, P. / Gao, X. / Cui, S. |
|---|
| 資金援助 |  中国, 1件 中国, 1件 | 組織 | 認可番号 | 国 |
|---|
| National Natural Science Foundation of China (NSFC) | 81772207 |  中国 中国 |
|
|---|
 引用 引用 |  ジャーナル: Acta Pharm Sin B / 年: 2022 ジャーナル: Acta Pharm Sin B / 年: 2022
タイトル: Acrylamide fragment inhibitors that induce unprecedented conformational distortions in enterovirus 71 3C and SARS-CoV-2 main protease.
著者: Qin, B. / Craven, G.B. / Hou, P. / Chesti, J. / Lu, X. / Child, E.S. / Morgan, R.M.L. / Niu, W. / Zhao, L. / Armstrong, A. / Mann, D.J. / Cui, S. |
|---|
| 履歴 | | 登録 | 2022年2月16日 | 登録サイト: PDBJ / 処理サイト: PDBJ |
|---|
| 改定 1.0 | 2022年6月22日 | Provider: repository / タイプ: Initial release |
|---|
| 改定 1.1 | 2022年6月29日 | Group: Database references / カテゴリ: citation
Item: _citation.journal_id_ISSN / _citation.pdbx_database_id_PubMed / _citation.title |
|---|
| 改定 1.2 | 2022年10月12日 | Group: Database references / カテゴリ: citation
Item: _citation.journal_volume / _citation.page_first / _citation.page_last |
|---|
| 改定 1.3 | 2023年11月29日 | Group: Data collection / Refinement description
カテゴリ: chem_comp_atom / chem_comp_bond / pdbx_initial_refinement_model |
|---|
| 改定 1.4 | 2025年9月17日 | Group: Advisory / Derived calculations / Structure summary
カテゴリ: pdbx_entry_details / pdbx_validate_close_contact / struct_conn
Item: _pdbx_entry_details.has_protein_modification |
|---|
|
|---|
 データを開く
データを開く 基本情報
基本情報 要素
要素 キーワード
キーワード 機能・相同性情報
機能・相同性情報
 X線回折 /
X線回折 /  シンクロトロン /
シンクロトロン /  分子置換 / 解像度: 2.05 Å
分子置換 / 解像度: 2.05 Å  データ登録者
データ登録者 中国, 1件
中国, 1件  引用
引用 ジャーナル: Acta Pharm Sin B / 年: 2022
ジャーナル: Acta Pharm Sin B / 年: 2022 構造の表示
構造の表示 Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク ダウンロード
ダウンロード 7wym.cif.gz
7wym.cif.gz PDBx/mmCIF形式
PDBx/mmCIF形式 pdb7wym.ent.gz
pdb7wym.ent.gz PDB形式
PDB形式 7wym.json.gz
7wym.json.gz PDBx/mmJSON形式
PDBx/mmJSON形式 その他のダウンロード
その他のダウンロード 7wym_validation.pdf.gz
7wym_validation.pdf.gz wwPDB検証レポート
wwPDB検証レポート 7wym_full_validation.pdf.gz
7wym_full_validation.pdf.gz 7wym_validation.xml.gz
7wym_validation.xml.gz 7wym_validation.cif.gz
7wym_validation.cif.gz https://data.pdbj.org/pub/pdb/validation_reports/wy/7wym
https://data.pdbj.org/pub/pdb/validation_reports/wy/7wym ftp://data.pdbj.org/pub/pdb/validation_reports/wy/7wym
ftp://data.pdbj.org/pub/pdb/validation_reports/wy/7wym リンク
リンク 集合体
集合体

 要素
要素

 X線回折 / 使用した結晶の数: 1
X線回折 / 使用した結晶の数: 1  試料調製
試料調製 シンクロトロン / サイト:
シンクロトロン / サイト:  SSRF
SSRF  / ビームライン: BL19U1 / 波長: 0.9789 Å
/ ビームライン: BL19U1 / 波長: 0.9789 Å 解析
解析 分子置換
分子置換 ムービー
ムービー コントローラー
コントローラー







 PDBj
PDBj