| Entry | Database: PDB / ID: 7w4b
|
|---|
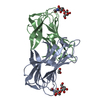
| Title | Phloem lectin (PP2) structure -complex with Chitotrise |
|---|
 Components Components | 17 kDa phloem lectin |
|---|
 Keywords Keywords | SUGAR BINDING PROTEIN / Chitin-binding protein / Phloem lectin |
|---|
| Function / homology | : / Phloem protein 2-like / Phloem protein 2 / carbohydrate binding / triacetyl-beta-chitotriose / 17 kDa phloem lectin Function and homology information Function and homology information |
|---|
| Biological species |  Cucumis sativus (cucumber) Cucumis sativus (cucumber) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.5 Å MOLECULAR REPLACEMENT / Resolution: 2.5 Å |
|---|
 Authors Authors | Sivaji, N. / Kishore, B.B. / Suguna, K. / Surolia, A. |
|---|
| Funding support |  India, 1items India, 1items | Organization | Grant number | Country |
|---|
| Department of Biotechnology (DBT, India) | |  India India |
|
|---|
 Citation Citation |  Journal: Structure / Year: 2023 Journal: Structure / Year: 2023
Title: Structure and interactions of the phloem lectin (phloem protein 2) Cus17 from Cucumis sativus.
Authors: Sivaji, N. / Kishore, B.B. / Suguna, K. / Surolia, A. |
|---|
| History | | Deposition | Nov 26, 2021 | Deposition site: PDBJ / Processing site: PDBJ |
|---|
| Revision 1.0 | Mar 8, 2023 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Nov 29, 2023 | Group: Data collection / Refinement description
Category: chem_comp_atom / chem_comp_bond / pdbx_initial_refinement_model |
|---|
| Revision 2.0 | May 22, 2024 | Group: Advisory / Atomic model ...Advisory / Atomic model / Data collection / Database references / Derived calculations / Other / Polymer sequence / Refinement description / Source and taxonomy / Structure summary
Category: atom_site / atom_sites ...atom_site / atom_sites / entity / entity_name_com / entity_poly / entity_poly_seq / entity_src_gen / pdbx_branch_scheme / pdbx_contact_author / pdbx_molecule_features / pdbx_poly_seq_scheme / pdbx_struct_assembly / pdbx_struct_assembly_gen / pdbx_struct_assembly_prop / pdbx_struct_sheet_hbond / pdbx_unobs_or_zero_occ_atoms / pdbx_unobs_or_zero_occ_residues / pdbx_validate_close_contact / pdbx_validate_torsion / refine / refine_hist / refine_ls_restr / refine_ls_restr_ncs / refine_ls_shell / reflns / software / struct_asym / struct_conf / struct_conn / struct_ncs_dom / struct_ncs_dom_lim / struct_ref / struct_ref_seq / struct_sheet_range
Item: _atom_sites.fract_transf_matrix[2][1] / _atom_sites.fract_transf_matrix[3][2] ..._atom_sites.fract_transf_matrix[2][1] / _atom_sites.fract_transf_matrix[3][2] / _entity.formula_weight / _entity.pdbx_description / _entity.pdbx_number_of_molecules / _entity_poly.pdbx_seq_one_letter_code / _entity_poly.pdbx_seq_one_letter_code_can / _entity_src_gen.gene_src_common_name / _entity_src_gen.pdbx_end_seq_num / _pdbx_struct_sheet_hbond.range_1_auth_comp_id / _pdbx_struct_sheet_hbond.range_1_auth_seq_id / _pdbx_struct_sheet_hbond.range_1_label_comp_id / _pdbx_struct_sheet_hbond.range_1_label_seq_id / _pdbx_struct_sheet_hbond.range_2_auth_comp_id / _pdbx_struct_sheet_hbond.range_2_auth_seq_id / _pdbx_struct_sheet_hbond.range_2_label_comp_id / _pdbx_struct_sheet_hbond.range_2_label_seq_id / _refine.B_iso_mean / _refine.aniso_B[1][1] / _refine.aniso_B[1][2] / _refine.aniso_B[1][3] / _refine.aniso_B[2][2] / _refine.aniso_B[2][3] / _refine.aniso_B[3][3] / _refine.correlation_coeff_Fo_to_Fc / _refine.correlation_coeff_Fo_to_Fc_free / _refine.details / _refine.ls_R_factor_R_free / _refine.ls_R_factor_R_work / _refine.ls_R_factor_all / _refine.ls_R_factor_obs / _refine.ls_d_res_low / _refine.ls_number_reflns_R_work / _refine.ls_number_reflns_obs / _refine.ls_percent_reflns_R_free / _refine.ls_percent_reflns_obs / _refine.ls_wR_factor_R_free / _refine.ls_wR_factor_R_work / _refine.overall_SU_B / _refine.overall_SU_ML / _refine.pdbx_R_Free_selection_details / _refine.pdbx_average_fsc_free / _refine.pdbx_average_fsc_work / _refine.pdbx_overall_ESU_R / _refine.pdbx_overall_ESU_R_Free / _refine.pdbx_stereochemistry_target_values / _refine.solvent_model_details / _refine_hist.cycle_id / _refine_hist.d_res_low / _refine_hist.number_atoms_total / _refine_hist.pdbx_number_atoms_ligand / _refine_hist.pdbx_number_atoms_protein / _reflns.number_obs / _software.version / _struct_conf.beg_label_seq_id / _struct_conf.end_label_seq_id / _struct_ref.pdbx_align_begin / _struct_ref.pdbx_seq_one_letter_code / _struct_ref_seq.db_align_beg / _struct_ref_seq.pdbx_auth_seq_align_beg / _struct_ref_seq.seq_align_end / _struct_sheet_range.beg_auth_comp_id / _struct_sheet_range.beg_auth_seq_id / _struct_sheet_range.beg_label_comp_id / _struct_sheet_range.beg_label_seq_id / _struct_sheet_range.end_auth_comp_id / _struct_sheet_range.end_auth_seq_id / _struct_sheet_range.end_label_comp_id / _struct_sheet_range.end_label_seq_id
Description: Ligand geometry / Provider: author / Type: Coordinate replacement |
|---|
| Revision 2.1 | Oct 16, 2024 | Group: Structure summary / Category: pdbx_entry_details / pdbx_modification_feature / Item: _pdbx_entry_details.has_protein_modification |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Cucumis sativus (cucumber)
Cucumis sativus (cucumber) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.5 Å
MOLECULAR REPLACEMENT / Resolution: 2.5 Å  Authors
Authors India, 1items
India, 1items  Citation
Citation Journal: Structure / Year: 2023
Journal: Structure / Year: 2023 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 7w4b.cif.gz
7w4b.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb7w4b.ent.gz
pdb7w4b.ent.gz PDB format
PDB format 7w4b.json.gz
7w4b.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/w4/7w4b
https://data.pdbj.org/pub/pdb/validation_reports/w4/7w4b ftp://data.pdbj.org/pub/pdb/validation_reports/w4/7w4b
ftp://data.pdbj.org/pub/pdb/validation_reports/w4/7w4b

 F&H Search
F&H Search Links
Links Assembly
Assembly

 Movie
Movie Controller
Controller



 PDBj
PDBj