[English] 日本語
 Yorodumi
Yorodumi- PDB-7u5a: Crystal structure of queuine salvage enzyme DUF2419 mutant K199C,... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7u5a | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
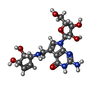
| Title | Crystal structure of queuine salvage enzyme DUF2419 mutant K199C, complexed with queuosine | |||||||||
 Components Components | Queuine salvage enzyme DUF2419 | |||||||||
 Keywords Keywords | HYDROLASE / 7-deazaguanine salvage / queuosine / tRNA modification | |||||||||
| Function / homology | Queuosine salvage protein family / Queuosine salvage protein / tRNA-guanine transglycosylation / Hydrolases; Glycosylases; Hydrolysing N-glycosyl compounds / hydrolase activity / MALONATE ION / Chem-QEO / Queuosine 5'-phosphate N-glycosylase/hydrolase Function and homology information Function and homology information | |||||||||
| Biological species |  Sphaerobacter thermophilus DSM 20745 (bacteria) Sphaerobacter thermophilus DSM 20745 (bacteria) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  FOURIER SYNTHESIS / Resolution: 2.5 Å FOURIER SYNTHESIS / Resolution: 2.5 Å | |||||||||
 Authors Authors | Hung, S.-H. / Swairjo, M.A. | |||||||||
| Funding support |  United States, 2items United States, 2items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res. / Year: 2023 Journal: Nucleic Acids Res. / Year: 2023Title: Structural basis of Qng1-mediated salvage of the micronutrient queuine from queuosine-5'-monophosphate as the biological substrate. Authors: Hung, S.H. / Elliott, G.I. / Ramkumar, T.R. / Burtnyak, L. / McGrenaghan, C.J. / Alkuzweny, S. / Quaiyum, S. / Iwata-Reuyl, D. / Pan, X. / Green, B.D. / Kelly, V.P. / de Crecy-Lagard, V. / Swairjo, M.A. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7u5a.cif.gz 7u5a.cif.gz | 283.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7u5a.ent.gz pdb7u5a.ent.gz | 229.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7u5a.json.gz 7u5a.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  7u5a_validation.pdf.gz 7u5a_validation.pdf.gz | 962.7 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  7u5a_full_validation.pdf.gz 7u5a_full_validation.pdf.gz | 968 KB | Display | |
| Data in XML |  7u5a_validation.xml.gz 7u5a_validation.xml.gz | 25.7 KB | Display | |
| Data in CIF |  7u5a_validation.cif.gz 7u5a_validation.cif.gz | 36.6 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/u5/7u5a https://data.pdbj.org/pub/pdb/validation_reports/u5/7u5a ftp://data.pdbj.org/pub/pdb/validation_reports/u5/7u5a ftp://data.pdbj.org/pub/pdb/validation_reports/u5/7u5a | HTTPS FTP |
-Related structure data
| Related structure data |  7u07SC  7u1oC  7u91C  7ugkC  7uk3C  7ulcC  8dl3C S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 37162.648 Da / Num. of mol.: 2 / Mutation: K199C Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Sphaerobacter thermophilus DSM 20745 (bacteria) Sphaerobacter thermophilus DSM 20745 (bacteria)Strain: DSM 20745 / S 6022 / Gene: Sthe_2331 / Production host:  #2: Chemical | #3: Chemical | ChemComp-MLI / | #4: Water | ChemComp-HOH / | Has ligand of interest | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.1 Å3/Da / Density % sol: 60.37 % / Description: Long, thin clustered crystals |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion / pH: 7 Details: 1.1 M Sodium malonate, 0.1 M HEPES, 0.5% (v/v) Jaffamine ED-2001 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL12-2 / Wavelength: 0.97946 Å / Beamline: BL12-2 / Wavelength: 0.97946 Å | ||||||||||||||||||||||||||||||
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jun 26, 2021 | ||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.97946 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.5→37.65 Å / Num. obs: 32478 / % possible obs: 100 % / Redundancy: 26.1 % / CC1/2: 0.998 / Rmerge(I) obs: 0.29 / Rpim(I) all: 0.057 / Rrim(I) all: 0.296 / Net I/σ(I): 12.8 | ||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  FOURIER SYNTHESIS FOURIER SYNTHESISStarting model: 7U07 Resolution: 2.5→37.65 Å / Cor.coef. Fo:Fc: 0.973 / Cor.coef. Fo:Fc free: 0.954 / WRfactor Rfree: 0.2017 / WRfactor Rwork: 0.1512 / FOM work R set: 0.8394 / SU B: 18.49 / SU ML: 0.167 / SU Rfree: 0.2155 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R Free: 0.216 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 148.81 Å2 / Biso mean: 54.753 Å2 / Biso min: 31.88 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.5→37.65 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.5→2.565 Å / Rfactor Rfree error: 0 / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller


 PDBj
PDBj