+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7fbr | ||||||
|---|---|---|---|---|---|---|---|
| Title | Solution structure of The first RNA binding domain of Matrin-3 | ||||||
 Components Components | Matrin-3 | ||||||
 Keywords Keywords | RNA BINDING PROTEIN / RRM / ALS/FTD / nuclear matrix protein / Structural Genomics / PSI-2 / Protein Structure Initiative / RIKEN Structural Genomics/Proteomics Initiative / RSGI | ||||||
| Function / homology |  Function and homology information Function and homology informationheart valve development / blastocyst formation / miRNA binding / ventricular septum development / post-transcriptional regulation of gene expression / activation of innate immune response / nuclear matrix / innate immune response / zinc ion binding / identical protein binding / nucleus Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR / simulated annealing | ||||||
 Authors Authors | Muto, Y. / Kobayashi, N. / Yokoyama, S. / RIKEN Structural Genomics/Proteomics Initiative (RSGI) | ||||||
 Citation Citation |  Journal: Biomol.Nmr Assign. / Year: 2022 Journal: Biomol.Nmr Assign. / Year: 2022Title: 1 H, 13 C and 15 N resonance assignments and solution structures of the two RRM domains of Matrin-3. Authors: He, F. / Kuwasako, K. / Takizawa, M. / Takahashi, M. / Tsuda, K. / Nagata, T. / Watanabe, S. / Tanaka, A. / Kobayashi, N. / Kigawa, T. / Guntert, P. / Shirouzu, M. / Yokoyama, S. / Muto, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7fbr.cif.gz 7fbr.cif.gz | 628.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7fbr.ent.gz pdb7fbr.ent.gz | 526.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7fbr.json.gz 7fbr.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  7fbr_validation.pdf.gz 7fbr_validation.pdf.gz | 431.6 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  7fbr_full_validation.pdf.gz 7fbr_full_validation.pdf.gz | 517.7 KB | Display | |
| Data in XML |  7fbr_validation.xml.gz 7fbr_validation.xml.gz | 30.5 KB | Display | |
| Data in CIF |  7fbr_validation.cif.gz 7fbr_validation.cif.gz | 53.4 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/fb/7fbr https://data.pdbj.org/pub/pdb/validation_reports/fb/7fbr ftp://data.pdbj.org/pub/pdb/validation_reports/fb/7fbr ftp://data.pdbj.org/pub/pdb/validation_reports/fb/7fbr | HTTPS FTP |
-Related structure data
| Related structure data |  7fbvC C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11351.972 Da / Num. of mol.: 1 / Mutation: S397R Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Production host: Cell-free gateway cloning vector N-term 8xHis eGFP pCellFree_G03 (others) References: UniProt: Q8K310 |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|---|
| NMR experiment | Sample state: isotropic / Type: 3D 13C,15N-SEPARATED NOESY SPECTRA |
- Sample preparation
Sample preparation
| Details | Type: solution Contents: 1.0 mM [U-99% 13C; U-99% 15N] RNA binding protein, 90% H2O/10% D2O Details: 1mM D-DTT;0.02% NaN3 were used to keep the protein condition Label: 13C,15N_sample / Solvent system: 90% H2O/10% D2O |
|---|---|
| Sample | Conc.: 1.0 mM / Component: RNA binding protein / Isotopic labeling: [U-99% 13C; U-99% 15N] |
| Sample conditions | Details: 20mM D-Tris-HCL (PH7.0); 100mM NaCl; 1mM D-DTT;0.02% NaN3 Ionic strength: 100 mM / Label: conditions_1 / pH: 7 / Pressure: 1 atm / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Bruker AVANCE / Manufacturer: Bruker / Model: AVANCE / Field strength: 800 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 1 Details: the structures are based on 2063 NOE-derived distance constrains, 41 main chain dihedral angle constraints based on TALOS program and 7 side chain dihedral constraints based on NOE pattern. | ||||||||||||||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with favorable non-bond energy Conformers calculated total number: 200 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller








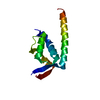


 PDBj
PDBj
 gel filtration
gel filtration