| 登録情報 | データベース: PDB / ID: 7fbl
|
|---|
| タイトル | Thrombocorticin in complex with Ca2+ and mannose |
|---|
 要素 要素 | Thrombocorticin |
|---|
 キーワード キーワード | SUGAR BINDING PROTEIN / bioactive protein / thrombocorticin |
|---|
| 機能・相同性 | alpha-D-mannopyranose 機能・相同性情報 機能・相同性情報 |
|---|
| 生物種 | Corticium sp. |
|---|
| 手法 |  X線回折 / X線回折 /  シンクロトロン / シンクロトロン /  分子置換 / 解像度: 1.419 Å 分子置換 / 解像度: 1.419 Å |
|---|
 データ登録者 データ登録者 | Kageyama, H. / Onodera, K. / Sakai, R. / Tanaka, Y. |
|---|
 引用 引用 |  ジャーナル: Nat Commun / 年: 2022 ジャーナル: Nat Commun / 年: 2022
タイトル: A marine sponge-derived lectin reveals hidden pathway for thrombopoietin receptor activation.
著者: Watari, H. / Kageyama, H. / Masubuchi, N. / Nakajima, H. / Onodera, K. / Focia, P.J. / Oshiro, T. / Matsui, T. / Kodera, Y. / Ogawa, T. / Yokoyama, T. / Hirayama, M. / Hori, K. / Freymann, D. ...著者: Watari, H. / Kageyama, H. / Masubuchi, N. / Nakajima, H. / Onodera, K. / Focia, P.J. / Oshiro, T. / Matsui, T. / Kodera, Y. / Ogawa, T. / Yokoyama, T. / Hirayama, M. / Hori, K. / Freymann, D.M. / Imai, M. / Komatsu, N. / Araki, M. / Tanaka, Y. / Sakai, R. |
|---|
| 履歴 | | 登録 | 2021年7月11日 | 登録サイト: PDBJ / 処理サイト: PDBJ |
|---|
| 改定 1.0 | 2022年7月13日 | Provider: repository / タイプ: Initial release |
|---|
| 改定 1.1 | 2023年7月19日 | Group: Data collection / Database references / カテゴリ: citation / citation_author / diffrn_source
Item: _citation.country / _citation.journal_abbrev ..._citation.country / _citation.journal_abbrev / _citation.journal_id_CSD / _citation.journal_id_ISSN / _citation.journal_volume / _citation.page_first / _citation.page_last / _citation.pdbx_database_id_DOI / _citation.pdbx_database_id_PubMed / _citation.title / _citation.year / _diffrn_source.pdbx_synchrotron_site |
|---|
| 改定 1.2 | 2023年11月29日 | Group: Data collection / Refinement description
カテゴリ: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / diffrn_source / pdbx_initial_refinement_model
Item: _diffrn_source.pdbx_synchrotron_site |
|---|
| 改定 1.3 | 2024年11月20日 | Group: Structure summary
カテゴリ: pdbx_entry_details / pdbx_modification_feature
Item: _pdbx_entry_details.has_protein_modification |
|---|
|
|---|
 データを開く
データを開く 基本情報
基本情報 要素
要素 キーワード
キーワード 機能・相同性情報
機能・相同性情報 X線回折 /
X線回折 /  シンクロトロン /
シンクロトロン /  分子置換 / 解像度: 1.419 Å
分子置換 / 解像度: 1.419 Å  データ登録者
データ登録者 引用
引用 ジャーナル: Nat Commun / 年: 2022
ジャーナル: Nat Commun / 年: 2022 構造の表示
構造の表示 Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク ダウンロード
ダウンロード 7fbl.cif.gz
7fbl.cif.gz PDBx/mmCIF形式
PDBx/mmCIF形式 pdb7fbl.ent.gz
pdb7fbl.ent.gz PDB形式
PDB形式 7fbl.json.gz
7fbl.json.gz PDBx/mmJSON形式
PDBx/mmJSON形式 その他のダウンロード
その他のダウンロード 7fbl_validation.pdf.gz
7fbl_validation.pdf.gz wwPDB検証レポート
wwPDB検証レポート 7fbl_full_validation.pdf.gz
7fbl_full_validation.pdf.gz 7fbl_validation.xml.gz
7fbl_validation.xml.gz 7fbl_validation.cif.gz
7fbl_validation.cif.gz https://data.pdbj.org/pub/pdb/validation_reports/fb/7fbl
https://data.pdbj.org/pub/pdb/validation_reports/fb/7fbl ftp://data.pdbj.org/pub/pdb/validation_reports/fb/7fbl
ftp://data.pdbj.org/pub/pdb/validation_reports/fb/7fbl リンク
リンク 集合体
集合体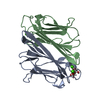
 要素
要素 Corticium sp. (in: Fungi) (菌類) / 発現宿主:
Corticium sp. (in: Fungi) (菌類) / 発現宿主: 
 X線回折 / 使用した結晶の数: 1
X線回折 / 使用した結晶の数: 1  試料調製
試料調製 シンクロトロン / サイト:
シンクロトロン / サイト:  Photon Factory
Photon Factory  / ビームライン: AR-NE3A / 波長: 1 Å
/ ビームライン: AR-NE3A / 波長: 1 Å 解析
解析 分子置換
分子置換 ムービー
ムービー コントローラー
コントローラー





 PDBj
PDBj







