| Entry | Database: PDB / ID: 6mib
|
|---|
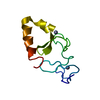
| Title | Crystal structure of the ILK ATP-binding deficient mutant (L207W)/alpha-parvin core complex |
|---|
 Components Components | - Alpha-parvin
- Integrin-linked protein kinase
|
|---|
 Keywords Keywords | CELL ADHESION / Pseudokinase / Protein complex |
|---|
| Function / homology |  Function and homology information Function and homology information
actin-mediated cell contraction / smooth muscle cell chemotaxis / establishment or maintenance of cell polarity regulating cell shape / protein localization to cell cortex / caveola assembly / Regulation of cytoskeletal remodeling and cell spreading by IPP complex components / Localization of the PINCH-ILK-PARVIN complex to focal adhesions / negative regulation of neural precursor cell proliferation / positive regulation of signal transduction / nerve development ...actin-mediated cell contraction / smooth muscle cell chemotaxis / establishment or maintenance of cell polarity regulating cell shape / protein localization to cell cortex / caveola assembly / Regulation of cytoskeletal remodeling and cell spreading by IPP complex components / Localization of the PINCH-ILK-PARVIN complex to focal adhesions / negative regulation of neural precursor cell proliferation / positive regulation of signal transduction / nerve development / fibroblast migration / establishment or maintenance of epithelial cell apical/basal polarity / myelination in peripheral nervous system / Cell-extracellular matrix interactions / positive regulation of BMP signaling pathway / outflow tract septum morphogenesis / sprouting angiogenesis / cell projection organization / neural precursor cell proliferation / heterotypic cell-cell adhesion / branching involved in ureteric bud morphogenesis / protein kinase inhibitor activity / establishment or maintenance of cell polarity / outflow tract morphogenesis / cilium assembly / positive regulation of osteoblast differentiation / positive regulation of substrate adhesion-dependent cell spreading / substrate adhesion-dependent cell spreading / cell-matrix adhesion / mitotic spindle organization / tumor necrosis factor-mediated signaling pathway / integrin-mediated signaling pathway / sarcomere / phosphatidylinositol 3-kinase/protein kinase B signal transduction / platelet aggregation / integrin binding / cell morphogenesis / Z disc / positive regulation of canonical Wnt signaling pathway / lamellipodium / actin cytoskeleton / actin binding / actin cytoskeleton organization / cell cortex / protein-macromolecule adaptor activity / cell differentiation / protein kinase activity / positive regulation of canonical NF-kappaB signal transduction / protein stabilization / cadherin binding / signaling receptor binding / focal adhesion / positive regulation of cell population proliferation / centrosome / protein kinase binding / positive regulation of DNA-templated transcription / chromatin / magnesium ion binding / nucleoplasm / ATP binding / membrane / nucleus / plasma membrane / cytoplasm / cytosolSimilarity search - Function Integrin-linked protein kinase, pseudokinase domain / Parvin / Calponin-like domain / Actin-binding Protein, T-fimbrin; domain 1 / Calponin homology domain / Calponin homology (CH) domain / : / Calponin homology domain / CH domain superfamily / Calponin homology (CH) domain profile. ...Integrin-linked protein kinase, pseudokinase domain / Parvin / Calponin-like domain / Actin-binding Protein, T-fimbrin; domain 1 / Calponin homology domain / Calponin homology (CH) domain / : / Calponin homology domain / CH domain superfamily / Calponin homology (CH) domain profile. / Ankyrin repeat profile. / Ankyrin repeats (3 copies) / Ankyrin repeat region circular profile. / ankyrin repeats / Ankyrin repeat / Ankyrin repeat-containing domain superfamily / Protein tyrosine and serine/threonine kinase / Serine-threonine/tyrosine-protein kinase, catalytic domain / Phosphorylase Kinase; domain 1 / Phosphorylase Kinase; domain 1 / Transferase(Phosphotransferase) domain 1 / Transferase(Phosphotransferase); domain 1 / Protein kinase domain profile. / Protein kinase domain / Protein kinase-like domain superfamily / 2-Layer Sandwich / Orthogonal Bundle / Mainly Alpha / Alpha BetaSimilarity search - Domain/homology |
|---|
| Biological species |  Homo sapiens (human) Homo sapiens (human) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å |
|---|
 Authors Authors | Fukuda, K. / Qin, J. |
|---|
| Funding support |  United States, 1items United States, 1items | Organization | Grant number | Country |
|---|
| National Institutes of Health/National Human Genome Research Institute (NIH/NHGRI) | |  United States United States |
|
|---|
 Citation Citation |  Journal: Nat Commun / Year: 2018 Journal: Nat Commun / Year: 2018
Title: Non-catalytic signaling by pseudokinase ILK for regulating cell adhesion.
Authors: Vaynberg, J. / Fukuda, K. / Lu, F. / Bialkowska, K. / Chen, Y. / Plow, E.F. / Qin, J. |
|---|
| History | | Deposition | Sep 19, 2018 | Deposition site: RCSB / Processing site: RCSB |
|---|
| Revision 1.0 | Nov 7, 2018 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Dec 18, 2019 | Group: Author supporting evidence / Category: pdbx_audit_support / Item: _pdbx_audit_support.funding_organization |
|---|
| Revision 1.2 | Mar 13, 2024 | Group: Data collection / Database references / Category: chem_comp_atom / chem_comp_bond / database_2
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession |
|---|
|
|---|
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å
MOLECULAR REPLACEMENT / Resolution: 1.8 Å  Authors
Authors United States, 1items
United States, 1items  Citation
Citation Journal: Nat Commun / Year: 2018
Journal: Nat Commun / Year: 2018 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 6mib.cif.gz
6mib.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb6mib.ent.gz
pdb6mib.ent.gz PDB format
PDB format 6mib.json.gz
6mib.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/mi/6mib
https://data.pdbj.org/pub/pdb/validation_reports/mi/6mib ftp://data.pdbj.org/pub/pdb/validation_reports/mi/6mib
ftp://data.pdbj.org/pub/pdb/validation_reports/mi/6mib Links
Links Assembly
Assembly
 Components
Components Homo sapiens (human) / Gene: ILK, ILK1, ILK2 / Production host:
Homo sapiens (human) / Gene: ILK, ILK1, ILK2 / Production host: 
 Homo sapiens (human) / Gene: PARVA, MXRA2 / Production host:
Homo sapiens (human) / Gene: PARVA, MXRA2 / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  APS
APS  / Beamline: 19-BM / Wavelength: 0.979 Å
/ Beamline: 19-BM / Wavelength: 0.979 Å Processing
Processing MOLECULAR REPLACEMENT / Resolution: 1.8→20.88 Å / Cor.coef. Fo:Fc: 0.961 / Cor.coef. Fo:Fc free: 0.937 / SU B: 2.186 / SU ML: 0.069 / Cross valid method: THROUGHOUT / ESU R: 0.111 / ESU R Free: 0.115 / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
MOLECULAR REPLACEMENT / Resolution: 1.8→20.88 Å / Cor.coef. Fo:Fc: 0.961 / Cor.coef. Fo:Fc free: 0.937 / SU B: 2.186 / SU ML: 0.069 / Cross valid method: THROUGHOUT / ESU R: 0.111 / ESU R Free: 0.115 / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS Movie
Movie Controller
Controller












 PDBj
PDBj