[English] 日本語
 Yorodumi
Yorodumi- PDB-6lvk: Crystal structure of FGFR2 in complex with 1,3,5-triazine derivative -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6lvk | ||||||
|---|---|---|---|---|---|---|---|
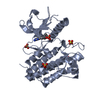
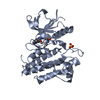
| Title | Crystal structure of FGFR2 in complex with 1,3,5-triazine derivative | ||||||
 Components Components | Fibroblast growth factor receptor 2 | ||||||
 Keywords Keywords | TRANSFERASE / protein kinase / SIGNALING PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationSignaling by FGFR2 amplification mutants / Signaling by FGFR2 fusions / fibroblast growth factor receptor signaling pathway involved in negative regulation of apoptotic process in bone marrow cell / fibroblast growth factor receptor signaling pathway involved in hemopoiesis / fibroblast growth factor receptor signaling pathway involved in positive regulation of cell proliferation in bone marrow / lateral sprouting from an epithelium / mammary gland bud formation / branch elongation involved in salivary gland morphogenesis / mesenchymal cell differentiation involved in lung development / lacrimal gland development ...Signaling by FGFR2 amplification mutants / Signaling by FGFR2 fusions / fibroblast growth factor receptor signaling pathway involved in negative regulation of apoptotic process in bone marrow cell / fibroblast growth factor receptor signaling pathway involved in hemopoiesis / fibroblast growth factor receptor signaling pathway involved in positive regulation of cell proliferation in bone marrow / lateral sprouting from an epithelium / mammary gland bud formation / branch elongation involved in salivary gland morphogenesis / mesenchymal cell differentiation involved in lung development / lacrimal gland development / prostate gland morphogenesis / otic vesicle formation / regulation of smooth muscle cell differentiation / regulation of morphogenesis of a branching structure / orbitofrontal cortex development / squamous basal epithelial stem cell differentiation involved in prostate gland acinus development / embryonic organ morphogenesis / branching morphogenesis of a nerve / morphogenesis of embryonic epithelium / endochondral bone growth / bud elongation involved in lung branching / epidermis morphogenesis / reproductive structure development / positive regulation of epithelial cell proliferation involved in lung morphogenesis / limb bud formation / gland morphogenesis / fibroblast growth factor receptor signaling pathway involved in orbitofrontal cortex development / ventricular zone neuroblast division / membranous septum morphogenesis / embryonic digestive tract morphogenesis / mesenchymal cell differentiation / positive regulation of phospholipase activity / epithelial cell proliferation involved in salivary gland morphogenesis / mesenchymal cell proliferation involved in lung development / FGFR2b ligand binding and activation / lung lobe morphogenesis / branching involved in prostate gland morphogenesis / branching involved in labyrinthine layer morphogenesis / FGFR2c ligand binding and activation / Activated point mutants of FGFR2 / Phospholipase C-mediated cascade; FGFR2 / regulation of osteoblast proliferation / fibroblast growth factor receptor activity / branching involved in salivary gland morphogenesis / embryonic cranial skeleton morphogenesis / lung-associated mesenchyme development / embryonic pattern specification / pyramidal neuron development / outflow tract septum morphogenesis / regulation of smoothened signaling pathway / mesodermal cell differentiation / bone morphogenesis / digestive tract development / positive regulation of mesenchymal cell proliferation / odontogenesis / ureteric bud development / organ growth / skeletal system morphogenesis / hair follicle morphogenesis / Signaling by FGFR2 IIIa TM / inner ear morphogenesis / lung alveolus development / prostate epithelial cord elongation / ventricular cardiac muscle tissue morphogenesis / regulation of osteoblast differentiation / prostate epithelial cord arborization involved in prostate glandular acinus morphogenesis / PI-3K cascade:FGFR2 / midbrain development / bone mineralization / positive regulation of cell division / fibroblast growth factor binding / fibroblast growth factor receptor signaling pathway / PI3K Cascade / epithelial to mesenchymal transition / negative regulation of keratinocyte proliferation / cell fate commitment / embryonic organ development / cellular response to transforming growth factor beta stimulus / regulation of ERK1 and ERK2 cascade / positive regulation of Wnt signaling pathway / positive regulation of cardiac muscle cell proliferation / SHC-mediated cascade:FGFR2 / positive regulation of vascular associated smooth muscle cell proliferation / cellular response to retinoic acid / FRS-mediated FGFR2 signaling / peptidyl-tyrosine phosphorylation / epithelial cell differentiation / positive regulation of cell cycle / Signaling by FGFR2 in disease / axonogenesis / lung development / excitatory synapse / positive regulation of epithelial cell proliferation / animal organ morphogenesis / post-embryonic development / Negative regulation of FGFR2 signaling / receptor protein-tyrosine kinase / bone development / Constitutive Signaling by Aberrant PI3K in Cancer / protein autophosphorylation Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.29 Å MOLECULAR REPLACEMENT / Resolution: 2.29 Å | ||||||
 Authors Authors | Echizen, Y. / Amano, Y. / Tateishi, Y. | ||||||
 Citation Citation |  Journal: Bioorg.Med.Chem. / Year: 2020 Journal: Bioorg.Med.Chem. / Year: 2020Title: Structure-based drug design of 1,3,5-triazine and pyrimidine derivatives as novel FGFR3 inhibitors with high selectivity over VEGFR2. Authors: Kuriwaki, I. / Kameda, M. / Hisamichi, H. / Kikuchi, S. / Iikubo, K. / Kawamoto, Y. / Moritomo, H. / Kondoh, Y. / Amano, Y. / Tateishi, Y. / Echizen, Y. / Iwai, Y. / Noda, A. / Tomiyama, H. ...Authors: Kuriwaki, I. / Kameda, M. / Hisamichi, H. / Kikuchi, S. / Iikubo, K. / Kawamoto, Y. / Moritomo, H. / Kondoh, Y. / Amano, Y. / Tateishi, Y. / Echizen, Y. / Iwai, Y. / Noda, A. / Tomiyama, H. / Suzuki, T. / Hirano, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6lvk.cif.gz 6lvk.cif.gz | 133.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6lvk.ent.gz pdb6lvk.ent.gz | 102.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6lvk.json.gz 6lvk.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lv/6lvk https://data.pdbj.org/pub/pdb/validation_reports/lv/6lvk ftp://data.pdbj.org/pub/pdb/validation_reports/lv/6lvk ftp://data.pdbj.org/pub/pdb/validation_reports/lv/6lvk | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6lvlC  6lvmC  3b2tS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 35886.215 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: FGFR2, BEK, KGFR, KSAM / Production host: Homo sapiens (human) / Gene: FGFR2, BEK, KGFR, KSAM / Production host:  References: UniProt: P21802, receptor protein-tyrosine kinase #2: Chemical | #3: Chemical | ChemComp-SO4 / #4: Water | ChemComp-HOH / | Has ligand of interest | Y | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.75 Å3/Da / Density % sol: 55.24 % |
|---|---|
| Crystal grow | Temperature: 296 K / Method: vapor diffusion, sitting drop / pH: 8.5 / Details: Tris, Ammonium sulfate, PEG 8000 |
-Data collection
| Diffraction | Mean temperature: 95 K / Serial crystal experiment: N | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU FR-E SUPERBRIGHT / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU FR-E SUPERBRIGHT / Wavelength: 1.5418 Å | ||||||||||||||||||||||||||||||
| Detector | Type: RIGAKU RAXIS VII / Detector: IMAGE PLATE / Date: Aug 30, 2010 | ||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.29→48.06 Å / Num. obs: 36222 / % possible obs: 99.7 % / Redundancy: 3.6 % / CC1/2: 0.997 / Rmerge(I) obs: 0.078 / Rpim(I) all: 0.047 / Rrim(I) all: 0.091 / Net I/σ(I): 9.7 | ||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3B2T Resolution: 2.29→48.06 Å / Cor.coef. Fo:Fc: 0.941 / Cor.coef. Fo:Fc free: 0.917 / SU B: 6.897 / SU ML: 0.164 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.267 / ESU R Free: 0.215 / Details: U VALUES : REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å | |||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 110.08 Å2 / Biso mean: 32.682 Å2 / Biso min: 11.79 Å2
| |||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.29→48.06 Å
| |||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.29→2.349 Å / Rfactor Rfree error: 0 / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller










 PDBj
PDBj









