Entry Database : PDB / ID : 6lplTitle A2AR crystallized in EROCOC17+4, SS-ROX at 100 K Adenosine receptor A2a,Soluble cytochrome b562,Adenosine receptor A2a Keywords / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Escherichia coli (E. coli)Method / / / Resolution : 2 Å Authors Ihara, K. / Hato, M. / Nakane, T. / Yamashita, K. / Kimura-Someya, T. / Hosaka, T. / Ishizuka-Katsura, Y. / Tanaka, R. / Tanaka, T. / Sugahara, M. ...Ihara, K. / Hato, M. / Nakane, T. / Yamashita, K. / Kimura-Someya, T. / Hosaka, T. / Ishizuka-Katsura, Y. / Tanaka, R. / Tanaka, T. / Sugahara, M. / Hirata, K. / Yamamoto, M. / Nureki, O. / Tono, K. / Nango, E. / Iwata, S. / Shirouzu, M. Funding support Organization Grant number Country Japan Society for the Promotion of Science (JSPS) JP16K14688
Journal : Sci Rep / Year : 2020Title : Isoprenoid-chained lipid EROCOC 17+4 : a new matrix for membrane protein crystallization and a crystal delivery medium in serial femtosecond crystallography.Authors: Ihara, K. / Hato, M. / Nakane, T. / Yamashita, K. / Kimura-Someya, T. / Hosaka, T. / Ishizuka-Katsura, Y. / Tanaka, R. / Tanaka, T. / Sugahara, M. / Hirata, K. / Yamamoto, M. / Nureki, O. / ... Authors : Ihara, K. / Hato, M. / Nakane, T. / Yamashita, K. / Kimura-Someya, T. / Hosaka, T. / Ishizuka-Katsura, Y. / Tanaka, R. / Tanaka, T. / Sugahara, M. / Hirata, K. / Yamamoto, M. / Nureki, O. / Tono, K. / Nango, E. / Iwata, S. / Shirouzu, M. History Deposition Jan 11, 2020 Deposition site / Processing site Revision 1.0 Nov 25, 2020 Provider / Type Revision 1.1 Nov 29, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accessionRevision 1.2 Nov 13, 2024 Group / Category / pdbx_modification_feature / Item
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human)
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å
MOLECULAR REPLACEMENT / Resolution: 2 Å  Authors
Authors Japan, 1items
Japan, 1items  Citation
Citation Journal: Sci Rep / Year: 2020
Journal: Sci Rep / Year: 2020 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 6lpl.cif.gz
6lpl.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb6lpl.ent.gz
pdb6lpl.ent.gz PDB format
PDB format 6lpl.json.gz
6lpl.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/lp/6lpl
https://data.pdbj.org/pub/pdb/validation_reports/lp/6lpl ftp://data.pdbj.org/pub/pdb/validation_reports/lp/6lpl
ftp://data.pdbj.org/pub/pdb/validation_reports/lp/6lpl


 10.5281/zenodo.3595784 / Data set type: diffraction image data
10.5281/zenodo.3595784 / Data set type: diffraction image data Links
Links Assembly
Assembly
 Components
Components Homo sapiens (human), (gene. exp.)
Homo sapiens (human), (gene. exp.) 

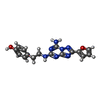






















 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SPring-8
SPring-8  / Beamline: BL32XU / Wavelength: 1 Å
/ Beamline: BL32XU / Wavelength: 1 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller













 PDBj
PDBj




























