[English] 日本語
 Yorodumi
Yorodumi- PDB-6l01: Crystal structure of E.coli DNA gyrase B in complex with 2-oxo-1,... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6l01 | ||||||
|---|---|---|---|---|---|---|---|
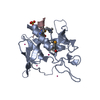
| Title | Crystal structure of E.coli DNA gyrase B in complex with 2-oxo-1,2-dihydroquinoline derivative | ||||||
 Components Components | DNA gyrase subunit B | ||||||
 Keywords Keywords | ISOMERASE / Inhibitor / Complex / Topoisomerase / Escherichia coli | ||||||
| Function / homology |  Function and homology information Function and homology informationAction of antimicrobials / DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) complex / DNA negative supercoiling activity / DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) activity / DNA topoisomerase (ATP-hydrolysing) / Antimicrobial resistance / DNA topological change / ATP-dependent activity, acting on DNA / DNA-templated DNA replication / chromosome ...Action of antimicrobials / DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) complex / DNA negative supercoiling activity / DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) activity / DNA topoisomerase (ATP-hydrolysing) / Antimicrobial resistance / DNA topological change / ATP-dependent activity, acting on DNA / DNA-templated DNA replication / chromosome / response to xenobiotic stimulus / response to antibiotic / DNA-templated transcription / DNA binding / ATP binding / metal ion binding / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.6 Å MOLECULAR REPLACEMENT / Resolution: 2.6 Å | ||||||
 Authors Authors | Mima, M. / Takeuchi, T. / Ushiyama, F. | ||||||
 Citation Citation |  Journal: Acs Omega / Year: 2020 Journal: Acs Omega / Year: 2020Title: Lead Identification of 8-(Methylamino)-2-oxo-1,2-dihydroquinoline Derivatives as DNA Gyrase Inhibitors: Hit-to-Lead Generation Involving Thermodynamic Evaluation. Authors: Ushiyama, F. / Amada, H. / Takeuchi, T. / Tanaka-Yamamoto, N. / Kanazawa, H. / Nakano, K. / Mima, M. / Masuko, A. / Takata, I. / Hitaka, K. / Iwamoto, K. / Sugiyama, H. / Ohtake, N. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6l01.cif.gz 6l01.cif.gz | 54.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6l01.ent.gz pdb6l01.ent.gz | 36.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6l01.json.gz 6l01.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/l0/6l01 https://data.pdbj.org/pub/pdb/validation_reports/l0/6l01 ftp://data.pdbj.org/pub/pdb/validation_reports/l0/6l01 ftp://data.pdbj.org/pub/pdb/validation_reports/l0/6l01 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6kzvC  6kzxC  6kzzC  1aj6S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 24191.182 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   References: UniProt: A0A4V5JMQ9, UniProt: P0AES6*PLUS, DNA topoisomerase (ATP-hydrolysing) |
|---|---|
| #2: Chemical | ChemComp-E0U / |
| #3: Water | ChemComp-HOH / |
| Has ligand of interest | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.97 Å3/Da / Density % sol: 37.47 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion / Details: MES, Ammonium acetate, PEG10000 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU MICROMAX-007 HF / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU MICROMAX-007 HF / Wavelength: 1.5418 Å |
| Detector | Type: RIGAKU RAXIS VII / Detector: IMAGE PLATE / Date: Dec 5, 2014 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.6→21.54 Å / Num. obs: 6243 / % possible obs: 100 % / Redundancy: 5.72 % / Rmerge(I) obs: 0.149 / Net I/σ(I): 7.8 |
| Reflection shell | Resolution: 2.6→2.69 Å / Rmerge(I) obs: 0.325 / Num. unique obs: 598 |
- Processing
Processing
| Software |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1AJ6 Resolution: 2.6→21.54 Å / Cross valid method: FREE R-VALUE
| ||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.6→21.54 Å
| ||||||||||||||||
| LS refinement shell | Resolution: 2.6→2.667 Å /
|
 Movie
Movie Controller
Controller











 PDBj
PDBj





