Entry Database : PDB / ID : 6b7kTitle GH43 Endo-Arabinanase from Bacillus licheniformis Endo-alpha-(1->5)-L-arabinanase Keywords / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / Biological species Bacillus licheniformis (bacteria)Method / / Resolution : 2.55 Å Authors Farro, E.G.S. / Nascimento, A.S. Funding support Organization Grant number Country Sao Paulo Research Foundation (FAPESP) 2014/03768-0 Sao Paulo Research Foundation (FAPESP) 2014/06565-2
Journal : Int. J. Biol. Macromol. / Year : 2018Title : GH43 endo-arabinanase from Bacillus licheniformis: Structure, activity and unexpected synergistic effect on cellulose enzymatic hydrolysis.Authors : Farro, E.G.S. / Leite, A.E.T. / Silva, I.A. / Filgueiras, J.G. / de Azevedo, E.R. / Polikarpov, I. / Nascimento, A.S. History Deposition Oct 4, 2017 Deposition site / Processing site Revision 1.0 Jun 13, 2018 Provider / Type Revision 1.1 Apr 17, 2019 Group / Data collection / Category / Item Revision 1.2 Jan 1, 2020 Group / Category / Item Revision 1.3 Oct 4, 2023 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Refinement description Category chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model / pdbx_struct_conn_angle / struct_conn Item _database_2.pdbx_DOI / _database_2.pdbx_database_accession ... _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_conn_angle.ptnr1_auth_asym_id / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr2_auth_asym_id / _pdbx_struct_conn_angle.ptnr2_auth_seq_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.55 Å
MOLECULAR REPLACEMENT / Resolution: 2.55 Å  Authors
Authors Brazil, 2items
Brazil, 2items  Citation
Citation Journal: Int. J. Biol. Macromol. / Year: 2018
Journal: Int. J. Biol. Macromol. / Year: 2018 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 6b7k.cif.gz
6b7k.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb6b7k.ent.gz
pdb6b7k.ent.gz PDB format
PDB format 6b7k.json.gz
6b7k.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/b7/6b7k
https://data.pdbj.org/pub/pdb/validation_reports/b7/6b7k ftp://data.pdbj.org/pub/pdb/validation_reports/b7/6b7k
ftp://data.pdbj.org/pub/pdb/validation_reports/b7/6b7k
 Links
Links Assembly
Assembly
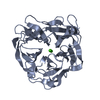



 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation ROTATING ANODE / Type: RIGAKU MICROMAX-007 / Wavelength: 1.5418 Å
ROTATING ANODE / Type: RIGAKU MICROMAX-007 / Wavelength: 1.5418 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller













 PDBj
PDBj

