+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5zt2 | |||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal structure of CCG DNA repeats at 1.66 angstrom resolution | |||||||||||||||||||||||
 Components Components | DNA (5'-D(* Keywords KeywordsDNA / DNA structures / i-motifs / neurological disease / tetraplex structures | Function / homology | : / DNA / DNA (> 10) |  Function and homology information Function and homology informationBiological species | synthetic construct (others) | Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 1.66002164937 Å MAD / Resolution: 1.66002164937 Å  Authors AuthorsHou, M.H. / Wu, P.C. / Satange, R.B. / Chen, Y.W. |  Citation Citation Journal: To Be Published Journal: To Be PublishedTitle: Crystallographic analysis of conformational change in CCG repeats into i-motif and unusual DNA duplex in presence and absence of CoII(Chro)2 complex Authors: Hou, M.H. #1:  Journal: Angew.Chem.Int.Ed.Engl. / Year: 2014 Journal: Angew.Chem.Int.Ed.Engl. / Year: 2014Title: Structural basis for the identification of an i-motif tetraplex core with a parallel-duplex junction as a structural motif in CCG triplet repeats. Authors: Chen, Y.W. / Jhan, C.R. / Neidle, S. / Hou, M.H. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5zt2.cif.gz 5zt2.cif.gz | 25.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5zt2.ent.gz pdb5zt2.ent.gz | 15 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5zt2.json.gz 5zt2.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/zt/5zt2 https://data.pdbj.org/pub/pdb/validation_reports/zt/5zt2 ftp://data.pdbj.org/pub/pdb/validation_reports/zt/5zt2 ftp://data.pdbj.org/pub/pdb/validation_reports/zt/5zt2 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
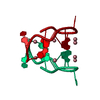
| Deposited unit | 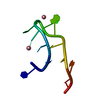
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| |||||||||||||||
| Unit cell |
| |||||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: DNA chain | Mass: 3295.150 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) | ||
|---|---|---|---|
| #2: Chemical | | #3: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.02 Å3/Da / Density % sol: 59.31 % |
|---|---|
| Crystal grow | Temperature: 277.15 K / Method: vapor diffusion, sitting drop / pH: 6 Details: 50mM Sodium cacodylate, 1mM Magnesium chloride, 5% MPD, 2mM Cobalt(II) chloride, 1mM Chromomycin A3 |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSRRC NSRRC  / Beamline: BL15A1 / Wavelength: 1.60572, 1.56518, 1.60490 / Beamline: BL15A1 / Wavelength: 1.60572, 1.56518, 1.60490 | ||||||||||||
| Detector | Type: RAYONIX MX-300 / Detector: CCD / Date: Oct 18, 2013 | ||||||||||||
| Radiation | Protocol: MAD / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||
| Radiation wavelength |
| ||||||||||||
| Reflection | Resolution: 1.66→30 Å / Num. obs: 3447 / % possible obs: 96.1 % / Redundancy: 5.5 % / Biso Wilson estimate: 16.5972142739 Å2 / Rmerge(I) obs: 0.048 / Net I/σ(I): 41.724 | ||||||||||||
| Reflection shell | Resolution: 1.66→1.72 Å / Rmerge(I) obs: 0.272 / Num. unique obs: 632 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MAD / Resolution: 1.66002164937→12.4437277752 Å / SU ML: 0.20873662372 / Cross valid method: FREE R-VALUE / σ(F): 0.00486031212139 / Phase error: 30.6258331204 MAD / Resolution: 1.66002164937→12.4437277752 Å / SU ML: 0.20873662372 / Cross valid method: FREE R-VALUE / σ(F): 0.00486031212139 / Phase error: 30.6258331204
| ||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||
| Displacement parameters | Biso mean: 26.7353127997 Å2 | ||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.66002164937→12.4437277752 Å
| ||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller









 PDBj
PDBj










































